Understanding viruses and challenges in microbiology
Issue: Why Microbiology Matters
05 May 2020 article
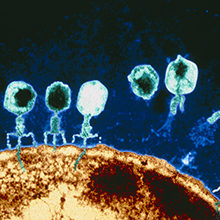
Virology holds a central position in both microbiology and public perception, never more than now as we face the challenge of a new viral pathogen. This section focuses on viruses, their structure and how we can manipulate viruses to benefit society.
Virology and viral disease
Nicola J. Stonehouse and Natalie Kingston
Viruses infect all forms of life, and while they can be extremely variable, the survival and propagation of all viruses is dependent upon living host cells. At their simplest, they are protein shells, with a nucleic acid centre. The shells (called capsids) protect the nucleic acid and serve to deliver this to new cells in order to spread infection. It is the nucleic acid that initiates disease. Cells can therefore become factories for the production of new viruses that then go on and infect other cells within the same host or infect new hosts.
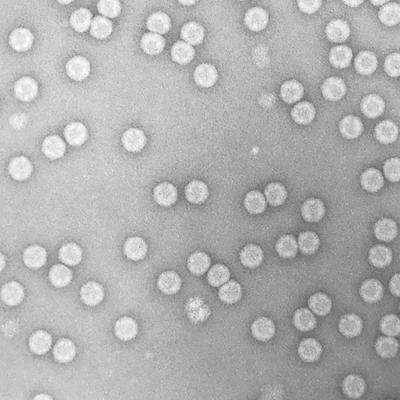
Although it is viruses such as Ebola and Zika that make the headlines, information from studies of a range of viruses is what informs the development of prophylactic vaccines and therapeutic treatments. Indeed, understanding such details of virus structure and lifecycle has revealed parallels in simple viruses that infect bacteria and yeast with those that infect plants and mammals. However, small changes can have big consequences in terms of both severity of infection and susceptibility of the host. Furthermore, new viruses are always emerging. This is mainly due to the speed at which the viral genomes are copied and the errors that can be introduced as a result. This ability to change quickly can mean that viruses can ’jump’ to infect new species.
Ongoing research is essential in order to better understand viruses, and to be in a position to respond rapidly to new and re-emerging viral disease.

Nicola J. Stonehouse
Faculty of Biological Sciences, University of Leeds, Leeds LS2 9JT, West Yorkshire, UK
[email protected]
@Stonehouse_Lab
Nicola Stonehouse was awarded a PhD in 1992 on dental enamel development. She then moved from studies of inorganic crystals to protein crystals and structural studies on RNA bacteriophages. Postdoctoral studies took her to Uppsala and Leeds and, in 1997, she was awarded an MRC Career Development Fellowship. Her research moved from phage to picornaviruses, maintaining a strong interest in RNA. She was appointed Lecturer in Leeds in 2001, then promoted to Senior Lecturer and to Chair of Molecular Virology in 2013.

Natalie Kingston
Faculty of Biological Sciences, University of Leeds, Leeds LS2 9JT, West Yorkshire, UK
[email protected]
@NatJKingston
Natalie Kingston completed a PhD at Monash University, Australia, in 2017. Her research focused on the generation of chimeric virus-like particle vaccines against Plasmodium. She then moved to the University of Leeds where she currently works on the characterisation of enteroviruses and the development of enterovirus vaccines with Nicola Stonehouse and David Rowlands.
What is the best career decision you’ve ever made?
Nicola: After my PhD, I moved fields to start working on viruses. At the time, this allowed me to translate my skills to a new biological problem, which was bacteriophage structure. But this opened up the world of viruses to me and I’m still fascinated by the virus lifecycle and of finding new antiviral strategies.
Natalie: Moving to Leeds and changing specialisation to molecular virology. This change has opened up new research areas for me that continue to be both exciting and have the potential to improve vaccine design.
Why does microbiology matter?
Nicola: Microbes are everywhere and affect almost all aspects of our lives. Harnessing the power of microbiology can therefore bring health, environmental, social, cultural, industrial and economic benefits to our society.
Viruses: the good, the bad and the useful
Hollie French, Elizaveta Elshina, Emmanuelle Pitre and Aartjan te Velthuis
Viruses are the most abundant and perhaps most diverse biological entities on Earth. They are simple life forms and are entirely dependent on hijacking host cells to replicate their genomes. However, contrary to common belief, not all viruses cause disease, since some are beneficial. By studying viruses, we can learn about the biology of host cells and organisms, develop strategies against viral disease and manipulate viruses for our own purposes.
Some viruses are only a single self-replicating gene, while others can encode almost a thousand proteins and be the size of a bacterium. Life cycles also vary among viruses, with some lasting millions of years and others less than an hour. Yet, in spite of vast structural and molecular differences, all viruses need to gain entry into a cell, find a site to replicate, and spread.
Binding and entry
Virus entry can only occur if a cell expresses surface proteins that a virus can bind to. Because this is not always the case, virus infections are restricted to specific cell types, organs and organisms. Knowledge of a virus’ tropism is important for estimating the potential that a virus emerges from a reservoir population and causes an outbreak in another species. For instance, a recently discovered bat influenza virus can enter human cells via the same receptor that it uses to enter bat cells, suggesting that bat flu might spill over into the human population.
Once inside a cell, viruses move their genetic material to sites of replication. Researchers can follow that movement by tracking single, fluorescently labelled viruses using a microscope. Studies of the interaction of the virus with the host have revealed how viruses use the cytoskeleton and intracellular membranes for viral translocation and replication, but also the importance of these host components for normal cellular signalling, movement and immunity.
Some viruses depend on the machinery in the nucleus and thus need to cross the nuclear membrane as well. To do this, influenza viruses and HIV-1 mimick host cell signals and employ cellular importins to carry their genome into the nucleus. In the nucleus, retroviruses express their genes by integrating them into the host genome. Such integration events can lead to cancer, but also shape animal evolution. One striking example is the ‘domestication’ of the retrovirus HERV-W envelope protein, now known as syncytin, as a component for placenta formation in mammals.
Replication and adaptation
Whether they replicate in the cytoplasm or nucleus, all viral genomes are copied by a polymerase. X-ray crystallography and electron microscopy techniques have enabled researchers to reveal the structure of these enzymes and develop drugs that frustrate viral replication. Some antiviral strategies can also exploit the high mutation rate of some viruses by pushing the error rate even higher, ultimately causing the viral population to collapse.
To prevent viruses from stealing their resources, cells express sensors that can identify viral genomes and proteins. The activation of these sensors triggers signalling, prevents virus spread and clears the infection. Many viruses encode proteins that can suppress or prevent these innate immune responses, but their function needs to be adapted to a host. Emerging viruses thus often trigger stronger responses than adapted viruses.
Vectors and spread
When new copies of the viral genome are ready to leave the host cell, some viruses fuse the infected cell with a neighbouring cell to allow faster spread. Other viruses condense their genome inside a protective protein shell with the right receptors to get the new virus to the next cell. To condense their genome, some herpesviruses and bacteriophages use a powerful molecular motor that can build up a pressure of 50 atmospheres!
Ultimately, a virus may need to spread between organisms. It can then rely on the natural behaviour of the host or a vector, such as a tick, or manipulate its host’s behaviour. The rabies virus, for instance, uses a snake-venom-like compound that makes animals aggressive and froth at the mouth with virus-laden saliva in order to increase the chance that a bite will spread the virus. Similarly, some baculoviruses can turn caterpillars into ‘zombies’ that climb up to high leaves and burst, spreading infectious virus particles to healthy caterpillars below.
Although viruses use their host, they are also incredibly useful. Molecular biology uses viral enzymes to manipulate RNA and DNA. Moreover, we can now alter viral receptors to re-target viruses to specific cells, such as cancerous cells. In addition, we can create attenuated viruses that can only proliferate in cancer cells, which often lack antiviral sensors; we can lyse tumours and keep healthy tissues, which are still able to control the infection, unharmed.
Viruses are masters at infecting cells, utilising life’s diverse abundance of molecules, systems and behaviours for their propagation. By studying them, we are learning from their expertise about ourselves and other organisms. This knowledge, combined with advances in other scientific fields, is enabling us to re-engineer viruses for our own purposes. Viruses may not only be the most abundant and diverse biological entities, but also some of the most useful.

Hollie French
Division of Virology, Department of Pathology, University of Cambridge, Cambridge CB2 0QQ, UK
[email protected]
@hollie_french20
Hollie French is a Research Assistant. She holds a BA in Natural Sciences (University of Cambridge) and now works on flu aberrant replication and innate immunity at the University of Cambridge. Her interests are in infectious disease virology and public health. She was previously an intern in the Global Polio Eradication Initiative (WHO, Geneva). Hollie has been a member of the Microbiology Society since 2017.

Elizaveta Elshina
Division of Virology, Department of Pathology, University of Cambridge, Cambridge CB2 0QQ, UK
[email protected]
@liza_elshina
After completing a BSc in Infectious Diseases at the University of Edinburgh, Elizaveta Elshina worked in preclinical vaccine development at the University of Oxford. She started to work on influenza virus during her MSc at the University of Zurich and is currently researching erroneous activity of influenza virus polymerase for her PhD. She has been a Microbiology Society member since 2018.

Emmanuelle Pitre
Division of Virology, Department of Pathology, University of Cambridge, Cambridge CB2 0QQ, UK
Emmanuelle Pitre graduated with a master’s in Fundamental Virology from Sorbonne University and the Pasteur Institute. She is now working towards a PhD at the University of Cambridge, on influenza viruses’ replication mechanisms.

Aartjan te Velthuis
Division of Virology, Department of Pathology, University of Cambridge, Cambridge, CB2 0QQ, UK
[email protected]
@aj_velthuis
lab.tevelthuis.com
Aartjan te Velthuis is a Henry Dale Fellow and Group Leader at the Department of Pathology of the University of Cambridge. He is interested in influenza virus replication and how aberrant viral RNA triggers innate immune responses. His research is funded by the Wellcome Trust, the Royal Society and NIH/NIAID.
Why does microbiology matter?
Joint answer: Microbiology has impacted human lives since the dawn of history. For centuries, it has played a key role in how we grow, prepare, flavour and preserve our foods. We used yeast for making beer before we knew how to make clean water and learnt to add salt to foods to prevent microbial growth. We also depend on microbiology to understand how viruses, bacteria and fungi cause disease, and how we can fight pathogens. In particular, the discovery of penicillin, a product of a fungus that can kill bacteria, has saved many lives. But microbiology is equally important for our future. It is helping us find ways to break down oil and plastic, to develop alternatives for other harmful products and to find (and possibly survive on) a habitable planet. Microbiology is without a doubt one of the most important research fields today.
What is the most rewarding part of your job?
Joint answer: In our lab, we study how influenza viruses replicate in human cells, how the viral genome is mutated, and how the efficiency of viral replication contributes to disease. It is extremely exciting to study this virus and being one of the first to uncover unknown molecular mechanisms. It feels like being an explorer discovering a new country or navigating a new mountain top. But one of the most rewarding aspects of our work is showing others how interesting microbiology is, either by presenting our work or using games such as our ‘virus roulette’ table to teach children (and adults) how infections and antibodies work.
Understanding viruses at the atomic scale: a history of virus structure research
David Bhella
As a PhD student in the crystallography department of Birkbeck University of London, I was struck by the proud heritage of that institution. Among the pioneers of structural biology from that department, J.D. Bernal, Rosalind Franklin and Aaron Klug made extraordinary contributions to structural virology. As a young researcher entering the field, a sense of walking in the footsteps of such towering historical figures was awe-inspiring. Over the intervening 25 years, I have been equally astounded by the technological developments in structural biology that have propelled the field forwards. In particular, the evolution of cryogenic electron microscopy (cryoEM), which has become a powerful tool for high-resolution structure determination, particularly suited to large macromolecular assemblies such as viruses.
Viruses are fascinating targets for structural biology research, being simultaneously the smallest (and most abundant) life-forms on the planet and the largest of macromolecular assemblies to be understood at the atomic level.
The shape of viruses
Our earliest insights into the shapes of viruses came when Helmut Ruska imaged plant and animal viruses for the first time in the transmission electron microscope (TEM). These images were published in 1939. At this time, J.D. Bernal and Isidor Fankuchen were beginning to work on X-ray diffraction of concentrated preparations of plant viruses, including tobacco mosaic virus (TMV) and tomato bushy-stunt virus (TBSV) – showing the former to be a long filamentous structure and the latter to be a spherical one.
The work started by Bernal was continued at Birkbeck college, where he recruited Rosalind Franklin to study the detailed structure of TMV. She showed it to be a helical assembly and defined the spatial arrangements of protein and RNA. Based on the work of Franklin’s collaborator and friend Don Caspar, Crick and Watson proposed that spherical viruses assemble with icosahedral symmetry. Using delightfully anachronistic language, these viruses were said to be likely to resemble a ‘rather symmetrical mulberry’ assembling from 60 protein subunits.
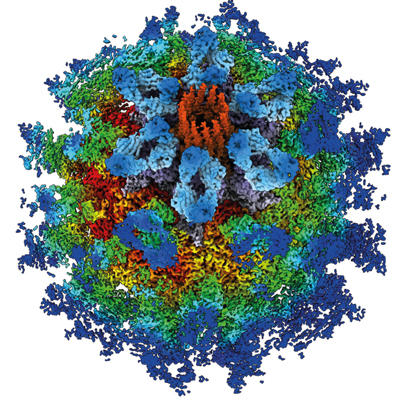
Initial theories of icosahedral symmetry in spherical viruses were insufficient, as many viruses were shown to form larger assemblies comprising many hundreds of capsid proteins. Don Caspar and Aaron Klug addressed this by formulating their theory of quasi-equivalent packing in icosahedral viruses. Building on emerging knowledge, they set out how larger capsids might be assembled by the introduction of small variations in bonding relationships between protein subunits.
Atomic modelling and structure
The first atomic model of a spherical virus was calculated for TBSV by Steve Harrison et al. in 1978, revealing a shell comprising 180 copies of the major capsid protein. This was followed by structures of two small RNA-containing viruses that infect humans: rhinovirus, solved by Michael Rossmann and colleagues, and poliovirus solved by Jim Hogle et al., both published in 1985. These studies revealed a common fold in the capsid proteins of these plant and animal viruses: an eight-stranded β-barrel known as the β-jelly roll.
The potential to use electrons rather than X-rays to determine virus structure was demonstrated in 1968 when David DeRosier and Aaron Klug calculated low-resolution 3D density maps of the contractile tail of phage T4 from TEM images. Exploiting the helical symmetry of the assembly allowed the density to be reconstructed from single images of phage particles stained with a heavy metal salt. A method to determine the structures of icosahedral objects followed, developed by Tony Crowther and colleagues. Many aspects of TEM were severely limiting, however, and the first step towards overcoming these challenges came with the invention of cryogenic methods for imaging biological material in the TEM. In 1985, Marc Adrian and colleagues published methods for the preparation of virus particles suspended in thin layers of vitreous ice. The absence of stain and chemical fixative meant that cryo-EM yielded images of macromolecular assemblies in a close to native state. Several technical advances in cryo-EM were required to move from early density maps at 30–40 angstroms resolution to where we are now – where 3D reconstructions at better than 4 angstroms resolution allow the construction of reliable atomic models.
Technological advances
The introduction of the first generation of digital cameras for TEM brought about a technological revolution in cryo-EM, facilitating the development of automated data collection and electron tomography (cryo-ET). Cryo-ET allows structure analysis of morphologically unique entities, by rotating them in the electron microscope and recording a tilt-series of images. These data may then be processed to compute a 3D reconstruction at intermediate resolution. A notable application of this method led to the calculation of an atomic model of the HIV capsid in the laboratory of John Briggs at EMBL Heidelberg.
For much of the first decade of the 21st century, virus structure research combined intermediate-resolution cryo-EM maps with X-ray data to yield pseudo-atomic models of, for example, complexes of viruses and host proteins such as antibodies. The first ab initio atomic model built into a single particle cryo-EM map of a virus was that of cytoplasmic polyhedrosis virus published by the laboratory of Z. Hong Zhou in 2008. The cryo-EM resolution revolution has since transformed this method to the point that atomic models of icosahedral virus capsids may be rapidly calculated. At the time of writing there are 175 capsid structures in the protein data bank solved by cryo-EM at better than 4 angstroms resolution. Notable recent achievements include high-resolution structures of two very large viruses: herpes simplex virus and African swine fever virus.
Recent developments in image reconstruction software have allowed investigators to probe asymmetry in viruses, revealing instances where deviating from symmetry is critical for the viral life cycle. One recent example from my own laboratory is our discovery that the calicivirus minor capsid protein VP2 forms a portal at a unique three-fold vertex following receptor binding. We believe that this is the mechanism by which the virus injects its genome into the cell.
Looking to the future of virus structure research, both X-rays and cryo-EM offer the tantalising prospect of viewing virus behaviour within the cell. Cryo soft X-ray microscopy is emerging as a powerful tool for imaging whole cells, revealing organelle rearrangements associated with virus infections. Cryo-ET of virus-infected cells is allowing researchers to analyse virus structures in situ, providing valuable biological context to structure data and promising that in the not too distant future it will be possible to solve structures of viruses in their natural habitat.
Harnessing structural biology in a crisis
In January 2020, SARS-Coronavirus 2 emerged in the Chinese city of Wuhan and has rapidly spread across the world, causing severe illness and deaths. This has led to the widespread lockdown of cities and whole nations. The scientific community has mobilised to address this crisis, including structural biologists. A testament to the significant advances in both X-ray crystallography and cryo-EM is the speed with which researchers have solved atomic structures for critical viral proteins. At the time of writing (20 March 2020), 29 protein structures for SARS-CoV-2 have been deposited in the Protein Data Bank, including the protease Mpro bound to several inhibitors, the S protein that mediates attachment and entry, and a complex of the S-protein’s receptor binding domain and the virus’ cellular receptor ACE2. These data will inform the development of antivirals and vaccines and are a major contribution to the global effort to defeat COVID-19.
David Bhella
MRC-University of Glasgow Centre for Virus Research (CVR), Sir Michael Stoker Building, Garscube Campus, 464 Bearsden Road, Glasgow G61 1QH, UK
[email protected]
@Davidbhella
David Bhella is Professor of Structural Virology at the Medical Research Council – University of Glasgow CVR. He is also Associate Director of the CVR and Director of the Scottish Centre for Macromolecular Imaging (SCMI). David started his career working as a diagnostic virologist at the Royal London Hospital before undertaking a PhD with Professor Helen Saibil FRS at Birkbeck College. He then moved to Glasgow’s MRC Virology Unit where he developed his programme of structural virology research.
Why does microbiology matter?
Microbiology impacts many key elements of human endeavour. Understanding microbes as primary drivers of the planet’s ecosystems, disease-causing agents, irreplaceable components of our own biological processes and as tools is a vital scientific need. Investigating the biology of micro-organisms has the potential to allow us to prevent and treat infectious and metabolic diseases, as well as feeding ourselves while minimising our ecological impacts.
What is the most rewarding part of your job?
For much of my career the primary driver has been the sheer thrill of discovery. That moment when your experiments lead to a new insight into the biology of an important virus is so often unexpected and startling. For a moment you are the only person in the history of humanity to know something important. Recent events have reminded me why I first chose to become a virologist. Understanding viruses at the molecular level is key to preventing serious diseases and saving lives. Microbiology is a worldwide endeavour and I am proud to play my own small part in this.
Images:
Thumbnail: Bacteriophage T4. Eye of Science/Science Photo Library.
Transmission electron micrograph of human rhinovirus, the main causative agent of the common cold. Nicola Stonehouse.
Asymmetry in icosahedral viruses – the calicivirus VP2 portal structure. https://www.ebi.ac.uk/pdbe/entry/emdb/EMD-0056.


