Schoolzone: Old School New School
09 August 2016
Determining phylogenetic relationships and evolutionary history of micro-organisms using bioinformatics
Bioinformatics is the method of using computer programming to interpret biological data. The teaching of bioinformatics in schools is becoming an integral part of the curriculum and understanding the evolutionary relationships between species at the genomic level is an important skill. However, the basic process of how these relationships are determined is fundamental in understanding all of these relationships.
The activities below demonstrate how the evolutionary history of different strains of a virus or other micro-organism can be determined by using both modern (computer) and historical (observation) methods.
PHYLOGENETIC COMPARISONS OF GENETIC SEQUENCE DATA TO DETERMINE THE ORIGINS OF HIV
HIV (human immunodeficiency virus) is a type of virus that attacks and weakens the immune system and most notably can lead to AIDS (acquired immune deficiency syndrome). Since the discovery of HIV in 1981, the origin of the virus has been widely debated. The most widely accepted hypothesis, which is backed up by the scientific evidence, is that HIV came from cross-species transmission of related viruses. Closely related non-human primates (monkeys and apes) carry viruses that are very similar to HIV, called SIVs (simian immunodeficiency viruses).
By comparing the genetic sequences of these closely related viruses it is possible to make assumptions about evolutionary relationships and origins of different strains. The following procedure is taken from a case study on the DNA to Darwin website. The activity makes use of Geneious software, which aligns protein sequences following nucleotide translation and builds phylogenetic trees that allow the user to visualise how closely related sequences are to one another.
Student procedure
- Download the free software, Geneious, from www.geneious.com and the Geneious document from www.dnadarwin.org/casestudies/7 containing the ‘DNA-HIV1andSIV.geneious’ file. This data file contains 16 nucleotide sequences of the pol gene from the HIV and SIV viruses from chimpanzees, gorillas and humans.
- Open the Geneious program and the DNA-HIV1andSIV.geneious file. The 16 sequences will be visible on the central window. Use the zoom function (magnifying glass button) to view the sequences at the nucleotide level. You should be able to see differences between the sequences at the base level. How can you tell the difference between an RNA and DNA virus at the nucleotide level?
- Make sure all 16 sequences are selected by clicking and highlighting the file name in the top window of the program. Click on the ‘Translate’ button to convert the nucleotide sequence to a protein sequence. (A box will appear, make sure the genetic code selected is ‘Standard’ and the translation frame is ‘1’).
- A new file will be created and as you did in Step 2, you can zoom in on this sequence and view the sequences at the amino acid level. Each letter represents a different amino acid. Can you guess which amino acid is represented by the letters?
- Now to ensure the sequences are aligned (they should be already, however). Make sure the protein sequence file is selected in the top window and select the ‘Alignment’ button. A box will appear and the ‘Geneious Alignment’ method needs to be selected (it is the only option with the basic software). The alignment can take a few minutes to process.
- A new alignment file will appear in the top window, select this and click on the ‘Tree’ button to create a phylogenetic tree. Another box will appear and the following options need to be selected; Genetic Distance Model: Jukes-Cantor, Tree build method: Neighbour-Joining, and Outgroup: No Outgroup. Make sure the ‘Resample Tree’ box is unchecked and select ‘OK’.
- Your phylogenetic tree should appear! You can zoom in and out and re-size the tree to properly study it. The tree should show you the different subgroups of the HIV-1 family and how these relate to the SIV sequences found in gorillas and chimpanzees. An easy way to visualise the tree is to print it out and highlight different relationships you can see.
Questions to consider
- Can you visualise the branches of the tree that are from human, chimpanzee or gorilla origin?
- How closely related are the different HIV subgroups to each other and to the SIV sequences. What can you conclude from this about the origins of HIV?
For further information, see our factfile on HIV and AIDS on our education website, Microbiology Online.
BUILDING A PHYLOGENETIC TREE BASED ON MORPHOLOGICAL CHARACTERISTICS
A simple way to make assumptions about the evolutionary history of a group of organisms is to observe morphological similarities between them and build a picture of how recently they shared a common ancestor based on the number of different features. The activity below is a basic introduction to how phylogenetic trees can be created using morphological characteristics of different micro-organisms. The trees will show common origins and how species can be related to one another based on morphology.
Student procedure
- Below are five illustrations (A–E) of well-known bacteria. They each have differences and similarities to one another.
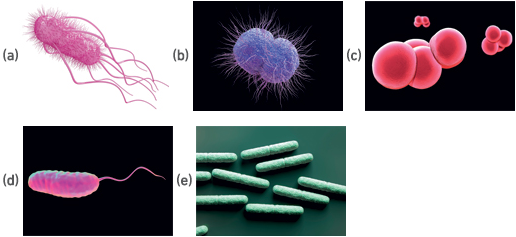
2. Using the table below, record the features for each of the micro-organisms. The features listed are purely suggestions, include your own.
- Using the table below, record the features for each of the micro-organisms. The features listed are purely suggestions, include your own.
|
|
Shape (rod or cocci) |
Colour |
Flagella present? |
Texture (smooth or spiked) |
|
A |
|
|
|
|
|
B |
|
|
|
|
|
C |
|
|
|
|
|
D |
|
|
|
|
|
E |
|
|
|
|
- In the following table, record the number of differences between each micro-organism. The numbers in each of the boxes indicate how the different micro-organisms relate to one another based on their morphology.
|
|
B |
C |
D |
E |
|
A |
|
|
|
|
|
B |
|
|
|
|
|
C |
|
|
|
|
|
D |
|
|
|
|
- To draw the phylogenetic tree, the five micro-organism letters will be placed at the top of the tree as these are the currently living micro-organisms. Firstly, you need to identify which of the micro-organisms are the closest relatives; these will have the least number of differences between them. Place this pair next to each other at the top of the tree. These two will share a recent common ancestor; place a dot below the two letters and draw lines leading to the letters showing their evolutionary lineages.
- To find the pairs of organisms that have a common ancestor that lived before the most recent pair, look for the micro-organisms that have two differences. Draw a dot below these organisms and link these too.
- Do the same for both three differences and when the organisms differ by four: this means they share the oldest common ancestor and the tree is complete.
An example of a smaller phylogenetic tree developed using this technique is shown below:
Things to consider:
- Are there other ways to classify the morphology of these micro-organisms? Think about possible lab techniques that you could use.
- Is it possible any of the morphological features have evolved independently of the other species (convergent evolution); are there ways you could determine this?
- Look at the world around you; think of other everyday items you could determine relatedness of with this technique. A tasty suggestion is a biscuit sampler tray; you could look at features such as chocolate coating and shape.
For further information about bioinformatics, see our open access journal, Microbial Genomics.
HANNAH FORREST
Public Affairs Administrator
[email protected]


