Microorganisms play a fundamental role in agriculture and food production, representing a key and indispensable resource that underpins the agri-food sector. Microbiota in these systems perform an array of pivotal functions essential to system health, sustainability and productivity. This conference will focus on the diverse roles played by microorganisms in agricultural systems, and on exploring what microbiome research can offer to agriculture. A particular emphasis will be placed on soil, plant and animal microbiomes, their interaction, and on cross-linking expertise across disciplines. An enhanced understanding of these microbiomes will provide opportunities towards managing agricultural systems in a manner that harnesses the natural power of microbes to provide solutions to global challenges of food security, resource limitation and climate change, and towards more efficient and sustainable food-production systems. This meeting will be of interest to academics, industry and government institutions.
The conference will incorporate a methods workshop that will enable integration of skills and sharing of best practice across technical aspects of microbiome research including sequencing and analysis/interpretation of data associated with complex microbial communities in agricultural systems. Leading experts in microbiome analysis will introduce novel technologies and give their perspectives on best practice in a series of invited talks and panel discussion. The methods workshop will be held at Teagasc (The Irish Agriculture and Food Development Authority), Moorepark in Fermoy, Co Cork. The conference will also include an optional tour of the genomic sequencing and microscopy facilities in Moorepark Research Centre.
This Focused Meeting will take place between 1-2 October 2018 at the Rochestown Park Hotel, Cork, Ireland. The methods workshop and optional site tour will be held on the afternoon of 2 October at Teagasc, Moorepark Research Centre, Fermoy, Co. Cork.
If you have any issues when attempting to register for this event, please contact [email protected]
Organising committee:
Key topics:
If you would like to register your interest or have an inquiry, please email [email protected]
Follow us on Twitter @MicrobioSoc, @soilmicrobio, @teagasc
Updates on Microbiomes Underpinning Agriculture can be found using the hashtag: #MUAFM18
Image: maxsattana/Thinkstock.
We are delighted to announce that the following speakers have been confirmed to speak at this Focused Meeting.

Dr Sharon Huws is a reader in Animal Science at the School of Biological Sciences and the Institute of Global Food Security, Queens University, Belfast. Her research is focused on understanding microbiomes, especially in the context of understanding the role that the rumen microbiome plays in ruminant food security. She is also interested in understanding the evolutionary drivers of antimicrobial resistance and exploiting microbiomes for industrial purposes. Sharon is also a senior editor for the journal ‘Microbiome’ and editor-in-chief for the new journal ‘Animal Microbiome. She also chairs the Rumen Microbial Genomics network, which underpins the activities of the Global Research Alliance. Dr Sharon Huws received her PhD from the University of Manchester, and subsequently went on to work as a post-doctoral scientist in the Universities of Bath and Aberystwyth. In 2010 she was promoted to Senior Principal investigator and later in 2012 was appointed as a lecturer, followed by progression to senior lecturer in 2015 at Aberystwyth University. She took up her new role in Queens University Belfast in 2017.

Davide Bulgarelli got his PhD in Plant Biology and Crop Production at the University of Milan (Italy). A postdoc position in the lab of Paul Schulze-Lefert at the MPI in Cologne (Germany) brought Davide in contact with the plant microbiota and it was “love at first sight…” In 2013 Davide moved to Scotland (UK), where he established his own lab and he is currently Principal Investigator at the Division of Plant Sciences, University of Dundee. Davide’s research focuses on understanding the genetic relationships between a plant genome and its associated microbiota and how these relationships can be exploited to sustainably enhance crop production.

Angela Sessitsch studied biochemistry at the University of Technology in Graz, holds a PhD in Microbiology from the Wageningen University, the Netherlands, and is habilitated at the Vienna University of Natural Resources and Life Sciences. She has pioneered plant-associated microbiomes, particularly in the endosphere, and she is interested in understanding the interactions between plants, microbiomes and the environment, as well as to develop applications. Her group explores the diversity and functioning of plant microbiota by applying a range of molecular approaches, interaction modes between plants and model bacteria, colonisation behaviour of endophytes, as well as various application technologies for biocontrol and crop enhancement applications. Together with her group, Angela has published more than 150 peer-reviewed publications.

Jana Seifert studied Geoecology at the Technical University of Freiberg, Germany. During her studies she got fascinated about microbiology and decided to move on this subject. She did her PhD in biology about the xenobiotic degradation by Gram-positive bacteria. As a postdoc she first focused on geomicrobiology and analysing microbial communities with molecular-genetic techniques. As a group leader at the Helmholtz Center for Environmental Research – UFZ in Leipzig, Germany she started to focus on the functionality of microbial communities setting up metaproteomics and protein-SIP. Since 2013, she is a junior professor at the University of Hohenheim, Germany, where she works on microbiome–animal interactions and the use of feeding resources by the gut microbiome.

After beginning his career as a chemist, with brief stints in the oil industry and NHS and a diversion into Mathematics, Leighton has worked his way up the scale through computational biology from small molecules, via snake venom toxins, to genomes. He was introduced to systems biology in his first two postdocs at Aberystwyth University, modelling yeast metabolism, and has been a computational biologist firstly at the Scottish Crop Research Institute, then the James Hutton Institute, since 2003. For the last 15 years Leighton has focused on biology at the interface of plants and their microbial pathogens, specialising in bacterial and oomycete pathogens of potato. Recently, he has been working on diagnostics and classification of microbial pathogens, and human pathogens in the environment.

Spencer Diamond is a microbiologist by training, but a bioinformatician by trade. He entered the scientific field during his time at the University of California Berkeley in 2005, where he worked in the lab of Prof. Louise Glass on the molecular mechanisms of cell fusion in the filamentous fungus Neurospora crassa. Subsequently in 2008 he began a second project within the newly created Energy Bioscience Institute at UCB to investigate N. crassa as a model for plant cell wall degradation and lignocellulosic ethanol production. After graduating in 2009 with a bachelors degree in Microbiology, Spencer undertook his PhD at the University of California San Diego, where he studied the control bacterial circadian rhythms exert over the control of cyanobacterial metabolism in the lab of Prof. Susan Golden. After graduating in 2016, he accepted a postdoctoral position back at UCB in the lab of Prof. Jill Banfield where he is working on large metagenomic datasets collected from soil to assemble underrepresented soil genomes and map their metabolic functions.

Orla is a Senior Computational Biologist in Teagasc Food Research Centre and a funded investigator with APC Microbiome Ireland and VistaMilk. Orla graduated from UCC with a BSc in Biochemistry and subsequently a PhD in Bioinformatics. As of August 2018 Orla has a H-Index of 43, having published over 80 peer-reviewed manuscripts, resulting in nearly 9100 citations. Her research focuses on elucidating the microbiome from various environments, including human gut and lung, rumen, food and soil.

Richard’s research is broadly concerned with understanding the role of interactions between plant and soil communities in regulating the structure and function of terrestrial ecosystems, and their response to global change. A particular goal of his research is to develop a mechanistic and conceptual understanding of how: (1) plant species and their traits influence soil biodiversity and ecosystem processes, such as carbon and nutrient cycling; (2) soil biodiversity influences nutrient cycling and plant community dynamics across different temporal and spatial scales; and (3) these interactions are affected by, and can potentially mitigate, climate change. A major focus of his current research is applying these concepts to the development of sustainable management options for agriculture, biodiversity and the delivery of ecosystem services, especially carbon sequestration and efficient nutrient cycling. Richard is currently President of the British Ecological Society and Senior Editor of Journal of Ecology.

Chris Creevey is a Professor of Computational Biology in the Institute for Global Food Security at Queen’s University Belfast. His group is interested understanding the complex interactions of natural microbial communities, using metagenomic, metatranscriptomic and metaproteomic data analyses. Novel computational approaches developed in his group utilise evolutionary and ecological concepts to understand the function and stability of natural host-associated microbial communities with a particular emphasis on the rumen microbiome. Chris received his Ph.D. in 2002 from the National University of Ireland (NUI) for his work in the area of phylogenetics and comparative genomics. Following this he worked as a postdoctoral researcher in NUI Maynooth Ireland and the European Molecular Biology Laboratory (EMBL) in Heidelberg, Germany. Before taking his current position, he held an SFI Stokes Lectureship in Teagasc, Ireland and a Readership in Aberystwyth University, Wales in Rumen Systems Biology.

Laurent Philippot is Director of Research at the French Institute for Agricultural Research (INRA) and is leading a research group at the department of Agroecology in Dijon. He did a sabbatical at Georgia Tech in Atlanta and at the Swedish University of Agricultural Science (SLU) in Uppsala in 2000 and 2009, respectively. His research focuses on bridging microbial community ecology, microbial processes and ecosystem functioning using microbial guilds involved in nitrogen cycling and greenhouse gas. He is serving as Senior Editor for The ISME Journal and as editorial board member for FEMS Microbiology Ecology and Applied and Environmental Microbiology. He has over 130 peer-reviewed articles, including Nature Climate Change, Nature Reviews Microbiology, The ISME Journal, Trends in Plant Science, Global Change Biology, etc. with ISI citations of >7000. He participated in several European research projects (Metaexplore, NORA, EcoFINDERS) and his currently involved in the ERA-NET Biodiversa project 'Digging Deeper'.

Christopher Quince (MRC Research Fellow and Associate Professor at the University of Warwick) is an internationally renowned expert in the bioinformatics and statistics of microbial community analysis using next generation sequence data. He develops novel algorithms that apply machine learning to the high throughput data generated by ‘omics technologies. He has published seminal highly-cited papers on processing 16S rRNA amplicons, in order to achieve accurate diversity estimates through the removal of sequencing errors and chimeras. Quince has also developed statistical methods for the probabilistic modelling of community structure. More recently he has focused on shotgun metagenomics, developing the CONCOCT and DESMAN algorithms to address the complex challenge of extracting species and strain genomes respectively from short read data.
Abstract submission is now closed.
All poster abstracts can be found in the attached abstract book:
In order to ensure your presentation runs smoothly, you are asked to comply with the following:
Those who are presenting a poster must ensure the work is presented as below. We cannot accommodate incorrectly formatted posters during the conference.
The Microbiology Society journal, Microbial Genomics, will be awarding a prize to the poster that presents particularly compelling or novel research in the field of genomics, including host–microbiota interactions and the microbiome. The winner will receive a cash prize and a certificate.
Registration is now open.
Please tick the option if you are planning to travel to Teagasc Moorepark by bus.
If you have any issues when attempting to register for this event, please contact [email protected]
| Registration categories |
|
Full price rate |
| Full and Full Concessionary Member (including dinner) | £200 | £210 |
| Full and Full Concessionary Member (excluding dinner) | £160 | £170 |
| Postgraduate and Undergraduate Student Member (including dinner) | £150 | £160 |
| Postgraduate and Undergraduate Student Member (excluding dinner) | £110 | £120 |
| Affiliate Member (including dinner) | £230 | £240 |
| Affiliate Member (excluding dinner) | £190 | £200 |
| Non-member (including dinner) | £260 | £270 |
| Non-member (excluding dinner) | £220 | £230 |
Upon registration you should receive an automated confirmation email. Please contact [email protected] if after 24 hours this has not been received.
If you need a letter of invitation for a visa application, we will be happy to supply this after we have received full payment.
The Department of Justice, Equality and Law Reform has primary responsibility for Ireland’s immigration and visa policy. For more information on visa requirements for Ireland please see their website: www.dfa.ie
Please note that all conference delegates are responsible for their own travel and visa arrangements; the Microbiology Society will not take any responsibility for travel or visa problems.
All registration fees must be paid in full BEFORE arrival at the meeting. Any outstanding registration fees must be paid before admittance will be granted to the meeting.
Refunds will not be provided.
Substitutions of attendees can be made at any time by contacting [email protected].
This meeting will take place at the Rochestown Park Hotel.
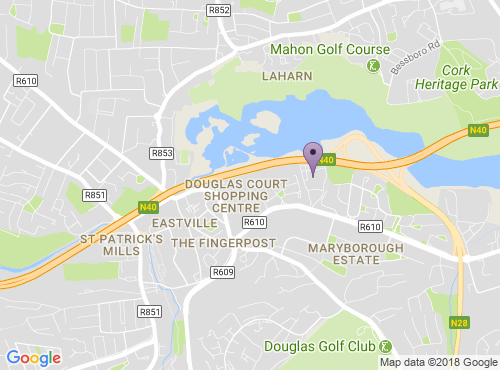
Rochestown Park Hotel,
Rochestown Road,
Douglas, Cork
Ireland
GPS Satellite Co-ordinates:
51.879425,-8.424863
Car parking
Complimentary parking is available for conference delegates.

Ireland’s second city is first in every important respect – at least according to the locals, who cheerfully refer to it as the ‘real capital of Ireland’. It’s a liberal, youthful and cosmopolitan place that was badly hit by economic recession but is now busily reinventing itself with spruced-up streets, revitalised stretches of waterfront, and – seemingly – an artisan coffee bar on every corner. There’s a developing hipster scene, but the best of the city is still happily traditional – snug pubs with live-music sessions, restaurants dishing up top-quality local produce, and a genuinely proud welcome from the locals.
The compact city centre is set on an island in the River Lee, surrounded by interesting waterways and packed with grand Georgian avenues, cramped 17th-century alleys and modern masterpieces such as the opera house.
St Patrick’s St runs from St Patrick’s Bridge on the North Channel of the Lee, through the city’s main shopping and commercial area, to the Georgian Grand Parade, which leads to the river’s South Channel. North and south of St Patrick’s St lie the city’s most entertaining quarters: grids of narrow streets crammed with pubs, shops, cafes and restaurants, fed by arguably the best foodie scene in the country.
Please visit Rochestown Park Hotel's website for further information.
The closest airport is Cork Airport, which is just over 7 kilometres (4.5 miles) away from Rochestown Park Hotel. There are no direct buses from the airport to the hotel, but the Air Coach Service (route 226) can take you to the city centre, where you can catch the 223 local bus to right outside the hotel (17 mins).
It is also possible to fly to Dublin Airport, then catch an Air Coach service to Cork Airport.
Find out more on Cork Airport's website.
By ferry
Cork Ringaskiddy Ferry Port is located 13.4 kilometres (8.3 miles) away from the venue (approx. 17 mins).
By train (within Ireland)
If travelling from Dublin: Trains from Dublin's Heuston rail station to Kent Station in Cork run regularly.
If travelling from Galway: There are no direct trains from Galway rail station to Kent Station, but there are connection trains.
Please see Iarnród Éireann (Irish Rail)'s website for timetables and more information.
By bus (within Ireland)
Please see Bus Éireann's website for more information.
We have been able to secure a discount on accommodation at the Rochestown Park Hotel. Please see the rates below, available for delegates attending our meeting in October.
Please use the following code for a discount: Micro18
Following the first two sessions of the conference, we would like to invite you to an informal drinks reception and poster session that will allow you to discuss the research with the authors and to catch up with old contacts and make new ones. Drinks will be served in the Kiltegan Suite from 5pm.
Drinks reception sponsored by APC Microbiome Ireland.
Conference dinner will be held in the Estuary Suite after the drinks reception. Three course dinner exclusively for the conference delegates will provide a great opportunity to continue the discussions from the day in a relaxed atmosphere.
Society Conference Grants will be available to support eligible members wishing to present at this Focused Meeting. Funding is also available for members requiring support for caring costs associated with conference attendance. The closing date for applications is Wednesday 12 September 2018. You can apply for a grant before receiving notification about the outcome of your abstract submission. A conditional grant offer can be made without evidence of abstract acceptance if unavailable at the time of application, however evidence must be provided to claim any grant offered.
Each year, the Young Microbiologist of the Year Competition recognises and rewards excellence in science communication by a Microbiology Society Member who is a postgraduate student or postdoctoral researcher, having gained their PhD in the last two years.
To enter this competition, applicants must tick the appropriate box during online abstract submission. Finalists will be invited to give a 10-minute oral presentation (plus 5 minutes for questions) at the final at the Society’s Annual General Meeting.
The ECM Forum Co-chairing Scheme provides ECM Forum members with the opportunity to be involved in the chairing of scientific sessions at the conference. The Co-Chairs will not receive any monetary value in co-chairing and will not take the place of a session Chair, but will receive a fantastic professional development opportunity to learn about being a session chair from more experienced colleagues.
ECM Forum members who are submitting an abstract to the meeting are asked to express interest in the Co-Chairing Scheme via the abstract submission system, and are invited to provide a statement outlining the following information:
All applications will be reviewed by the Society's Divisions and successful Co-Chairs will be introduced to the relevant session Chair.
Co-Chairs will receive a letter of thanks from the ECM Forum Executive Committee confirming that they participated in the Co-Chairing Scheme, and will be recognised in the conference programme.
For questions about the ECM Forum Co-chairing Scheme, please contact [email protected].
Exhibition and sponsorship opportunities are available for this meeting. For more information, please contact [email protected] or download the exhibition pack.

|

|

|

|

|

|

|