Microbiology and genetics
Issue: Why Microbiology Matters
05 May 2020 article
The genetic plasticity of microbes enables their survival. This section outlines some of the advances that have been made in modern biotechnology and the significance of this research to microbiology.
Genetics in microbiology
Charles J. Dorman
Much of our understanding of mutation, genome rearrangement, DNA repair, DNA recombination mechanisms, DNA replication, DNA transfer, gene regulation and much more has come from fundamental research in microbes. Their short life cycles have allowed microbes to be used in studies of evolution that have elucidated many of the underlying molecular processes. During World War Two, studies in a bacterium (Pneumococcus) identified DNA as the agent that could transform the phenotype of a living cell. Over sixty years ago, the first genetic switches to be characterised were in bacteria, and we continue to learn about bacterial gene regulation mechanisms and about genome-wide, collective control of gene expression. This both informs our understanding of naturally occurring microbes and guides developments in synthetic biology. Microbial and viral genetic switches are exploited as logic gates in biological computers, blurring boundaries between biology and information technology. Simultaneously, advances in conventional computing power allow us to interpret unprecedented volumes of genetic information, facilitating new approaches to infectious disease surveillance, prediction, prevention and therapy.
|
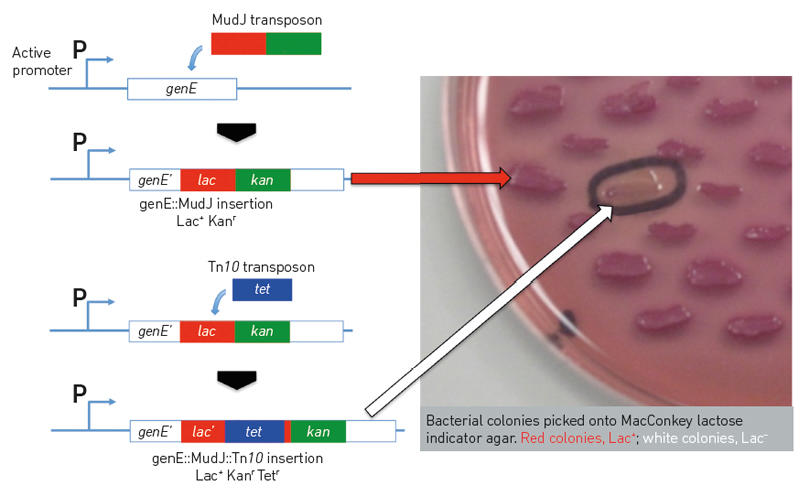
| A simple experiment in bacterial genetics exploiting ‘jumping genes’ to create gene fusions and genetic mutations. A gene of interest in Salmonella, genE, is tagged with a lac reporter gene using a modified Mu bacteriophage , MudJ. Insertion of the modified phage by transposition is selected on kanamycin-containing agar plates followed by screening for the presence of the desired genE-lac fusion on MacConkey lactose plates where these form red (Lac+) colonies. The genE promoter, P, is responsible for expression of the Lac+ phenotype and any signals affecting its activity can now be studied in living bacteria that harbour the genE-lac fusion. The lac reporter gene is next interrupted by insertion of a Tn10 transposon, producing colonies that are white (Lac–) on MacConkey lactose indicator plates. This proves that the lac gene is required for the red Lac+ phenotype. The structures of the modified genes are confirmed by polymerase chain reaction amplification followed by DNA sequencing. Charles J. Dorman |
Modern biotechnology is founded on microbial systems. Recombinant DNA methodology (‘cloning’) began with bacterial plasmid vectors into which foreign DNA was introduced deliberately. Cloning exploits bacterial restriction enzymes, proteins that ‘restrict’ the replication of bacteriophages (viruses) in bacteria. Bacterium-specific DNA methylation patterns are epigenetic features that protect their genomes from restriction. The polymerase chain reaction (PCR) is based on the heat-resistant polymerases found in archaea from high-temperature environments. PCR is now in routine use in molecular biology and in forensic laboratories. The public has become accustomed to descriptions in the news media of CRISPR-Cas as a ‘gene editing tool’. In fact, it is an adaptive immunity system used by bacteria to identify and to destroy invading DNA molecules, including viruses.
Combining microbial genetics with single-molecule and single-cell techniques, together with advances in imaging such as super resolution microscopy, is providing understanding of molecular processes at unprecedented levels of detail. The future for genetics in microbiology is a very bright one indeed.
Further reading
Dorman CJ. Structure and Function of the Bacterial Genome. Hoboken, NY, USA: John Wiley and Sons; 2020.

Charles J. Dorman
Trinity College Dublin, The University of Dublin, College Green, Dublin 2, Ireland
[email protected]
Charles J. Dorman joined the Microbiology Society in 1982. He has held the established Chair of Microbiology at Trinity College Dublin in Ireland since 1994, having previously been a lecturer in biochemistry at Dundee University in Scotland, where he also held a Royal Society University Research Fellowship.
Why does microbiology matter?
Studying the relationships that microbes have with one another, with non-microbial organisms and with the wider environment provides humans with useful insights into how these transactions can be conducted successfully and sustainably. They may be much smaller than us, but microbes face problems of resource discovery and management, energy deficits, transport and manufacturing, waste production and recycling, political interactions involving cooperation and antagonism, economic cycles of boom-and-bust and much more that is eerily reminiscent of our own experiences.
What inspired you to become a microbiologist?
Two things. (1) Discovering that a study of bacteria opened the door to an investigation of gene regulation in biological systems that are experimentally tractable. (2) Discovering an undergraduate degree course that allowed me to win enough prize money each year to cover the following year’s tuition fees (unlike the UK in the 1970s and early 1980s, Ireland charged for tuition, making university education difficult to access for people from my social group).
Restriction enzymes
Qaiser I. Sheikh and David P. Hornby
Restriction enzymes, or endonucleases, have been at the core of molecular biology for over 50 years. This is perhaps somewhat surprising, in view of the esoteric nature of the work that led up to the discovery of these enzymes, which was famously recognised by the award of the 1978 Nobel Prize in Medicine or Physiology to Werner Arber, Daniel Nathans and Hamilton O. Smith. The practical translation of an enzymatic system, referred to both genetically and functionally as ‘restriction and modification’, represents a landmark in the close interplay between biological genome processing and the laboratory manipulation of genes. In many prokaryotes, restriction and modification form part of a struggle for supremacy between bacteria and bacteriophages. The restriction enzyme attacks and destroys unmodified DNA (the bacteriophage) while the modification enzyme protects the host genome from restriction.
Restriction enzymes have been centre stage in molecular biology for around 40 years, providing the first route to molecular cloning of genes in prokaryotes. The type II restriction enzymes (often considered by aficionados to be the boring relations in this fascinating family of endonucleases, which are categorised with respect to their genetic and biochemical properties) are among the most enduring and valuable tools for molecular biologists. These homo-dimeric enzymes typically recognise a short (4–8 base pairs) DNA duplex within a genome or plasmid and introduce a precise cut across the symmetry axis, producing either two ‘blunt’ DNA fragments, in a sequence such as that shown below:

or a staggered cut, in a palindromic sequence of the type shown below:

In the laboratory, any ‘compatible’ DNA fragments can be joined using, for example, a DNA ligase enzyme. Restriction enzymes are given names that reflect their biological origin and the chronology of their discovery, hence EcoRI (from Escherichia coli strain R, recognising the duplex GAATTC) and BamHI (from Bacillus amyloliquefaciens strain H, recognising the palindromic sequence GGATCC). Restriction and modification enzymes play key roles in safeguarding the host genome in many microbial genomes, deriving their name from their function in restricting the growth of invading bacteriophages.
The golden age of restriction enzymes
During the ‘golden age’ of restriction enzymes in molecular biology, they were essential for not only cutting and pasting genes, but were central to the development of DNA diagnostics – some of you may remember RFLPs (restriction fragment length polymorphisms) and for the construction of plasmid and later genome maps. Ten years after the first commercial development of restriction enzymes, pioneered by New England Biolabs in the late 1970s, the emergence of thermostable DNA polymerases took centre stage, as the polymerase chain reaction began to drive the development of many molecular biology protocols. However, there remain today many situations, such as analytical gene and plasmid mapping and the general manipulation of DNA fragments, that enable these two powerful technologies to exist side by side.
The first steps to CRISPR-Cas9
In a similarly serendipitous discovery, a series of clustered, regularly interspaced short palindromic repeats discovered in prokaryotes 15 years later led to the development of the contemporary field of gene and genome editing. Unlike restriction enzymes, the recognition sequences are derived from prior phage infection and genome recombination events. However, the first step in the CRISPR-Cas9 (in one well used example) process is a sequence-specific DNA cleavage, in a manner related to the endonuclease activity characteristic of restriction enzymes. Biologically, this is used to fend off subsequent phage invasion by the original phage, but the ability to harness this phenomenon for any specific gene has led to its emergence as one of the most important techniques in genomic medicine as well as molecular biology research.
Restriction (and modification) enzymes have not only powered biological and medical research and diagnostics since their discovery, they have also shed light on the fundamental molecular events associated with DNA recognition. Starting with the publication of the structure of EcoRI and EcoRV in the 1990s, the diversity of recognition strategies, including DNA twisting, kinking and flipping (discovered first in the modification partner of HhaI restriction enzyme), have paved the way for our molecular understanding of many genetic transactions. Moreover, the growth of microbial genome sequences has revealed that restriction and modification genes are present in around 90% of those genomes sequenced to date. In my view, this fascinating class of genes has many more secrets to reveal.

Qaiser I. Sheikh
The Krebs Institute, Department of Molecular Biology and Biotechnology, University of Sheffield, Firth Court, Western Bank, Sheffield S10 2TN, UK
Qaiser I. Sheikh studied in Pakistan and Japan before moving to the University of Sheffield for a PhD and postdoctoral research fellowships. He then worked as Postgraduate Support Officer and then as a lecturer in the Department of Molecular Biology and Biotechnology, and developed the current Molecular Biology and Biotechnology MSc course, of which he is Programme Leader.

David P. Hornby
The Krebs Institute, Department of Molecular Biology and Biotechnology, University of Sheffield, Firth Court, Western Bank, Sheffield S10 2TN, UK
[email protected]
+44 (0)114 222 4232
David P. Hornby obtained a BSc (Biochemistry and Chemistry) and a PhD (Biochemistry) from the University of Sheffield. He undertook postdoctoral work in Sheffield, Leeds and Basle, followed by a Fellowship at EMBL and work at the Scripps Research Institute. He was appointed to a Lectureship at Sheffield in 1990 and became Director of Enterprise in Life Sciences in 2008. He was Sabbatical Visitor (2012–2013) at the LSTM and Director of Research and Innovation at the Liverpool Life Science UTC (2013–2015). He is currently working with school students in association with the Northern Schools Trust.
Why does microbiology matter?
David: I used to place a high value on microbiology because it made biochemistry more tractable: a somewhat simplistic and rather selfish perspective, but this was because I spent much of my research career purifying and characterising enzymes and later recombinant enzymes. Many years later, as I develop new multidisciplinary science lab classes for schools based around insects and simple catalysts, I have become fascinated by the concept of the microbiome and the evolution of early microbes. Microbiology matters to me now because it helps us not only understand how life originated on Earth, but how microbes continue to play a key role in the evolution of all higher organisms through a diverse range of mutually beneficial interactions.
What is the most rewarding part of your job?
David: Being able to continually educate myself and then finding a way to pass the ideas and concepts on to young scientists of all abilities.
CRISPR-Cas: Cunning Research Investigating the Science of Prokaryotes Results in Challenges and Applications for Society
Peter C. Fineran and Nils Birkholz
It has been a common theme in recent years: just before the Nobel Prize winners are announced, the #CRISPR (clustered regularly interspaced short palindromic repeats) Twitter feeds are buzzing with debates. Which heroes and/or heroines of the field will be awarded? October 2019 was no different. And even though all expectations went unfulfilled once more, everyone agrees that a CRISPR Nobel is a question of when not whether. Who should the Academy pick, and for what? The characterisation of CRISPR as a sophisticated form of bacterial immunity? The discovery that the enzyme Cas9, involved in this immunity, could be harnessed for gene editing purposes? One of the many CRISPR-based technologies developed since? The impact of CRISPR is not only founded in the tremendous research output, however. If, within a decade, a scientific discovery rises to a global issue worthy of dedicated documentaries, the impact of this discovery is going to be profound – not only for academia but for society as a whole. How did it come to this?
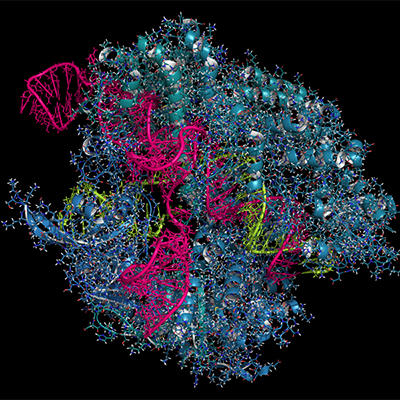
CRISPRs and Cas
It is a beautiful example of the serendipity of fundamental research leading to unexpected potential in diverse applications. It all started with some peculiar repetitive sequences in a bacterial genome, more than 30 years ago. It took decades to figure out that these sequences, now known as CRISPRs, provide an immune memory – storing small bits of DNA from past invaders, such as viruses, to facilitate their recognition upon subsequent attack. Scientists from all over the planet have now elucidated more details of how bacteria update and use these memories, as well as CRISPR-associated (Cas) proteins, to detect and destroy their enemies. Cas9 attracted particular attention after it was shown to be a single programmable nuclease – it can be supplied with an artificial memory and target any desired sequence which will then be cleaved. The researchers envisioned that this capacity would enable the application of Cas9 for gene editing.
The use of CRISPR-Cas
Bacteria possessing an adaptive immune mechanism – a feature long thought to be exclusive to higher organisms – was a spectacular discovery in its own right, but the easy programmability of Cas9 really got the ‘CRISPR Craze’ rolling. Not long after, CRISPR technologies were applied in fields such as agriculture, where plants can be enhanced to withstand drought or increase yield, and diverse model and previously ‘non-model’ organisms are being genetically modified with ease. The possibilities of the technology appear endless. Why not apply it to human genomes to rid them of disease-causing mutations? Indeed, multiple clinical trials involving medical applications of genome editing are being pursued.
This is exciting – but CRISPR–Cas9 is not flawless. For example, Cas9 not only targets the programmed sequence but can sometimes cause damage to other genomic locations too. To make precise changes in the genome, researchers can supply a repair template, but this is not always efficient. Newer variants of the technology aim to address these challenges. One approach is to use a Cas9 variant that can target but not cleave DNA, and couple this with another protein that performs a certain task at the desired location. For example, base editors can alter a single DNA base without cutting the double helix – useful, as many genetic diseases are caused by a single faulty nucleotide. The recently developed technique of prime editing utilises Cas9 fused to a reverse transcriptase enzyme for a find-and-replace approach.
Ethics and prohibitions
While we are getting closer to mastering the technical challenges of gene editing, many other aspects require consideration before it can be widely applied. With the rapid development of CRISPR technologies, ethics, public acceptance and legislation have a hard time catching up. A striking demonstration of this was the announcement in late 2018 by a Chinese scientist who claimed to have used Cas9-mediated genome editing on twin embryos to protect them against HIV infection. Although many countries prohibit human germline modifications, the outrage that followed the announcement highlighted the lack of adequate controls and regulations. Some leading scientists have called for a moratorium, while others consider this a waste of time or question the feasibility: it would be difficult to monitor and not all countries might care equally about prohibitions.
Reaching consensus is paramount, not only for genome editing but for other CRISPR-based innovations too, as the latter might be jeopardised if scepticism against CRISPR arises. For example, Cas9 and other CRISPR-associated proteins, such as Cas12 and Cas13, can be utilised for the detection of viral infections or identifying antibiotic-resistant bacteria – important applications in times of devastating viral epidemics and failure of common antibiotics. Things need to be handled transparently for CRISPR technologies to unfold to their full potential. Yet more innovations are certain to be discovered – and with them, yet more discussion for the yearly Twitter Nobel debates.

Peter C. Fineran
Department of Microbiology and Immunology, University of Otago, PO Box 56, Dunedin 9054, New Zealand
[email protected]
@PeterFineran
Peter C. Fineran heads a research team interested in bacterial defence systems, including CRISPR-Cas and bacteriophage biology. He obtained his PhD from the University of Cambridge, UK, and a BSc (Hons) in Biochemistry from the University of Canterbury, NZ. He has been a Microbiology Society member since 2002 and was the recipient of the 2019 Fleming Prize.

Nils Birkholz
Department of Microbiology and Immunology, University of Otago, PO Box 56, Dunedin 9054, New Zealand.
[email protected]
@NilsBirkholz
Nils Birkholz is a PhD candidate working on interactions between bacteria and bacteriophages in Peter Fineran’s research group. He obtained his BSc and MSc degrees from the Technical University of Braunschweig, Germany.
How does CRISPR benefit the average person?
Peter: Well, if you had yoghurt this morning then you probably had some dairy that has been fermented using strains that have been naturally adapted (using CRISPR) to phages that can spoil fermentations in the dairy industry. So, it is already having an application there. More generally, the tools are leading to massive advances in our ability to do fundamental science and to understand disease processes. We are also heading in the direction of being able to use CRISPR in different therapies and diagnostics.
Why does microbiology matter?
Peter: Although invisible, microbes are everywhere and important for so many things, from the environment, to health and industry. For me, I love that microbiology is such an incredible experimental system. That is actually why I got interested in microbial genetics; it’s kind of like solving a puzzle. You can just play around with the components and then actually work out how systems work. For example, you can just take out a gene, see what happens; put a gene in, see what happens. It’s because of this simplicity that microbiology is such a powerful system for answering biological questions.
Image: CRISPR-CAS9 gene editing complex from Streptococcus pyogenes. The Cas9 nuclease protein (blue) uses a guide RNA (green) sequence to cut DNA (pink) at a complementary site. Molekuul/iStock.
