Unlocking the world of microbiomes: exploring microbial communities
Issue: Why Microbiology Matters
05 May 2020 article

Research into the microbiome has evolved over time, allowing us to study microbial communities, genes and proteins in more depth. This section considers some of the interactions between microbial life and their environments in this developing area of science.
Microbial communities
Lindsay Hall
We live in a microbial world. Rather than living in isolation, the vast majority of microbes reside within communities, i.e. microbiomes, which play crucial roles in their ‘home’ environment. Thus, as microbiologists we are in a unique position to explore and seek answers to many different global challenges. We can look inside us to understand how our resident microbes impact our health and also look out to the microbes that influence the food that we eat and the environments that animals and crops inhabit. We can study the environmental microbes that reside within the places we live, work and play; at a global level we can focus on how microbes modulate our climate and the planet that we call home; and even further afield, to boldly go where no microbiologist has gone before. Focusing on how bacteria, viruses, fungi, archaea and microscopic eukaryotes interact with each other, their niche and their ‘host’ has opened up new avenues for microbiologists to explore, innovate and have impact in. The microbiome field has welcomed with open arms other disciplines within science and the humanities, as well as those working within healthcare, industry, policy and agriculture settings, together with the wider public, whose thirst for knowledge on microbiomes seems insatiable. Like our microbial friends, scientists (and microbiologists!) do not live in isolation, but rather thrive within supportive and dynamic communities. It’s been 75 years since the Microbiology Society was founded, with some key ‘microbiome’ discoveries made throughout this period driving real impacts, like the introduction of faecal microbiota transplants to treat Clostridioides difficile infection. Here’s to the next 75, and further exciting encounters within a world of microbiomes.

Lindsay Hall
Quadram Institute Bioscience, Norwich Research Park, Norwich, Norfolk NR4 7UQ, UK
Lindsay Hall is a Microbiome Group Leader and Wellcome Trust Investigator. Her team study the early life gut microbiota–host interactions, with a particular focus on Bifidobacterium. Lindsay obtained a BSc in Microbiology from the University of Glasgow, then a PhD in Microbiology and Immunology from the University of Cambridge and was a Postdoctoral Fellow at University College Cork, Ireland, before returning to the UK to work at the University of East Anglia and the Quadram Institute Bioscience from 2015.
Why does microbiology matter?
Microbes are all around us – and play hugely important roles in all aspects of our lives. It’s pretty amazing that things so small have such significant impacts, and therefore why being a microbiologist is so exciting, as we can really strive to make a difference.
Please tell us a little about the education and outreach activities you undertake in your role.
As well as our lab and clinical work, the team is also passionate about wider public engagement. One of our larger projects involved working with artists and creators to develop a giant walkthrough interactive gut to convey the importance of our gut microbes. We have also more recently developed an exciting new primary school resource which is aimed at introducing children to their ‘Guardians of the Gut’, a project which has been supported by the Microbiology Society.
Understanding and harnessing the wild microbiome
Jack A. Gilbert
When and why did I become known as a ‘microbiome researcher’? To be honest I cannot remember when, but it was probably because ‘microbiome’ became the new zeitgeist that helped us to get funding. I have always considered myself a microbial ecologist, studying the world of microbe–microbe interactions, and how these are shaped by the environment, be that an ocean, soil or human gut. Microbiome research is relatively field agnostic; for example, I currently reside in the Department of Pediatrics and Scripps Institution of Oceanography at University of California, San Diego. I guess that makes me the world’s first paediatric oceanographer, or oceanographic paediatrician? Unfortunately, neither of these epithets is supported by my CV, but they do extol the breadth of my research interests. Of course, I would consider my research interests quite narrow – I really only study how microbes interact within a given ecosystem, and how that shapes who grows and who dies – competition, mutualism, parasitism and commensalism; but I have the opportunity to apply that to any environment, including human bodies, plant roots, industrial factories and even buildings.
Industrial microbiology

Historically, the study of microbes has been dominated by those that cause disease, but if we strip away that 50,000 lb gorilla, we are left with the study of microbes of utility to industry and exploration of their role in environmental ecology. Industry-focused research has been dominated by microbial fermentation of foods for preservation, food spoilage and agriculture – with attempts to harness microbial chemistry in industrial processes relegated to the Holy Grail bin. Mind you, that is not to say that we have not been able to harness microbial chemistry in industrial processes, just that the number of successes is greatly outweighed by the number of failures, much like science generally, I suppose. The basic premise for discovery of novel functionalities across all industrial and agricultural practice has been to look for microbially mediated processes in natural systems, including extreme natural environments, food and soils, and attempts to isolate those organisms that perform the observed activities. For example, the discovery of yeasts as agents of fermentation, demonstrated by Pasteur (actually the work of many different scientists over many decades), led to the isolation of yeasts and the development of an industry using yeasts and their products, which is still flourishing today. The sheer number of ways in which microbial chemistry has been harnessed for industrial processes is not worth listing here, but it is exciting to note that new techniques are being derived all the time to augment our microbial toolbox. The application of metagenomic and metatranscriptomic sequencing, for example, is being used to mine the genomes of microbes across the planet for important enzymes – the diversity of coding both at the nucleic acid and amino acid level means that new variants of enzymes and chemical transformation pathways can now be catalogued, synthesised in the lab, transformed into heterologous expression models and even integrated into a synthetic organism and used to augment an existing chemical strategy. Coming up with new ways of producing nutrients in the lab, or improving the productivity, disease resistance and stress resilience of crops and animals, could help bolster food security with a growing population and the threat of climate change. Work to capture those microbes that grow on our pollution (e.g. oil spills, plastics, chemical dump sites) and harness them to clean up our mess is also a critical need. These applied approaches require further funding to tackle the worst pollution problems facing our planet – including carbon dioxide and methane production: identifying microbes that can store or transform these pollutants would be transformative.
Environmental microbiome
Since the father of modern ecology, G. Evelyn Hutchinson, identified the ooze at the centre of his dynamic food webs, we have been interested in how microbes shape our visible ecosystems. The history of environmental microbiology is long and detailed, and in many ways, while ostensibly a basic science with no obvious translational impact, it has influenced the industrial and medical microbiology fields far more than most people expect. Demonstrating that most microbes in most environments are centrally important to the proper functioning of an ecosystem helped to instill in people the concept that non-pathogenic microbial metabolism may be important in agricultural and clinical settings. With the ground-breaking technical advances pioneered by Carl Woese and the intellectual contributions from scientists like Jo Handlesman, we have launched environmental microbiome research into a new era. My own contribution, the Earth Microbiome Project (EMP), provided the impetus to leverage techniques developed through pioneering efforts from Julie Huber and Mitch Sogin to catalogue the world’s microbial diversity. The study, which is in fact a massive-super collaboration, has now generated data on more than 100,000 environmental samples and an equal number of human samples. It has demonstrated the ubiquity of microbial distribution, supporting the preposition that everything has the potential to be everywhere, but the environment selects for traits, resulting in environment-specific adaptations that can also propagate throughout the global ecosystem. The concept of a global ‘seed bank’ which was proposed by Jay Lennon in 2011, and further supported by the EMP in 2017, creates a framework for modelling and predicting how the Earth microbiome will respond to a changing climate and land use. That’s the next frontier, leveraging the ever-growing body of data to allow us to better steward the planet by giving us forewarning of impending doom, as well as potentially the tools to address it.
Biobanking for the future
Of course, our efforts to use microbes in industry and agriculture, as well as the potential to manipulate the environmental microbiome to effect changes in the equilibrium that could benefit our survival, may be moot if we cannot access the necessary microbial diversity. That is why myself and my colleagues are building a vision to install a microbiome seed bank in non-sovereign territory that could hold examples of the Earth’s microbiome. Guided by the data from the EMP, we are identifying samples of whole ecosystems, including soils, water, air, plants, animals, etc. that can be stored in a viable but dormant state, with the aim of resurrecting these genomic resources when we need them most. The Microbiota Vault will be a repository of the planet’s microbial heritage that could be our saviour if we are unable to halt global destruction.

Jack A. Gilbert
Department of Pediatrics and Scripps Institution of Oceanography, University of California San Diego School of Medicine, San Diego, 9500 Gilman Dr, La Jolla, CA 92093, USA
[email protected]
@gilbertjacka
Jack Gilbert is a Professor of Paediatrics. He helped establish the Microbiota Vault and works on the American Gut Project and Earth Microbiome Project. He began his career as an ecologist at the Natural History Museum in London before undertaking his PhD at Nottingham University, a postdoctoral position at Queen’s University in Ontario, Canada, and a directorship at the University of Chicago’s Microbiome Center working on the Home Microbiome Project and Hospital Microbiome Project before moving to San Diego in 2019.
Why does microbiology matter?
Quite simply, microbiology is the study of the lifeblood of our planet. If we can fathom the depth of knowledge that microbes possess, we could reshape medicine, revolutionise farming and industry, colonise barren worlds and hopefully limit the damage we are inflicting on our own planet.
What do you love most about your job?
Diversity. I love that I get to work with so many unique and brilliant people and engage on a plethora of wonderful projects. Working on microbial ecology, I can be at the bottom of the ocean, in the beating heart of a hospital or on a rocket to mars. Nothing is off limits.
Microbiomes underpinning agricultural systems
Sharon Huws
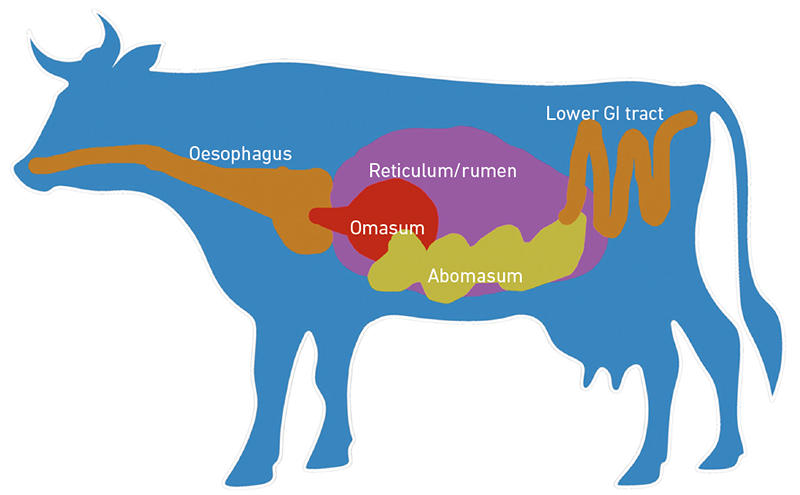
Microbes associated with crops, livestock and agricultural environments are essential for food security and food safety. For example, microbes in and around plant root systems, the rhizosphere, play a role in plant growth, nutrient absorption, disease resistance and soil structure. Microbes associated with food production, storage and transport can be implicated in food spoilage and the spread of pathogens. Livestock gastrointestinal tract microbes also contribute substantially to host production, health and environmental impact of livestock agriculture. In particular, the rumen microbiome is known to influence feed efficiency, quality of ruminant products and the extent of nitrogen and methane released to the environment (Fig. 1). Indeed, the influence of microbes on agricultural systems and food production has been known for many years.
Microbiomes and ‘omic technologies
The advent and rapid growth of ‘omic technologies in the past few years have increased our understanding of microbe diversity and function and have led to the terminology ‘microbiome’ being developed to describe microbial diversity and function in a biome. Arguably the most used terminology in the world of microbiology in the past decade is the term ‘microbiome’. The Nobel Laureate, Professor Joshua Lederberg, is often quoted as being the person to coin the term in 2001 to describe a collection of microbes living within a biome. Nonetheless, a small number of publications pre-dating this use the term ‘microbiome’ to denote the same meaning. Irrespective, how has the coining of this term underpinned by developments in ‘omic technologies helped us advance science? For many years post the advent of the first nucleic acid sequencers, studies on agriculture-related microbiomes, as well as microbiomes in general, were largely focused on metataxonomy-based studies, i.e. sequencing rRNA to assess microbial diversity within a biome. Monitoring diversity alone can be argued as being akin to stamp-collecting as it does not give us information on the function of the microbiome. Nonetheless, metataxonomical approaches are cheaper than function-based ‘omic technologies and software is now available allowing a broad inference of function from 16S rDNA information. The lack of standardisation and use of controls in these studies in the past have also meant that comparing between studies has been challenging due to the number of variables in protocols. This also leads to our inability to come to conclusions on a global level. Consequently, it could be argued that few of these metataxonomic studies have moved the field on substantially, although baseline information of the underlying microbial diversity in these biomes has aided our understanding of the ‘core’ microbes, which are always found in these biomes. One such study is the global rumen census study, which provided metataxonomic information for rumen-faecal microbiomes from 768 samples taken from a range of ruminants and has allowed the definition of the ‘core’ rumen/faecal microbes across the various geographical locations.
More recently published articles using multi-omics (metagenomics/metatranscriptomics/metaproteomics/metabolomics) and system-biology-based approaches have provided a major step-change in our understanding of microbiome function. For example, these approaches aid on-farm antimicrobial resistance (AMR) risk management strategies and define target genes/proteins for animal production/health/environmental impact purposes. These multi-omic approaches also allow us to prospect for useful compounds and enzymes in agriculture-based biomes, for further characterisation and commercial development. For example, a number of novel glycosyl hydrolases (carbohydrate-active enzymes) have been discovered within livestock gastrointestinal tract microbiomes which have a plethora of industrial uses, e.g. bioethanol production.
|
| Fig. 1. Ruminant gastrointestinal tract. Most of the microbes (bacteria, archaea, fungi, protozoa and phage) inhabit the large fermentative rumen, resulting in effective digestion of dietary material. |
Metagenomically-assembled genomes and culturomics
Sequencing technologies were largely used to circumvent the challenges associated with isolating representative bacteria using culture-based techniques, the so called ‘great plate anomaly’.
However, until recently it was not possible to isolate and construct genomes from metagenomes, which meant that assigning functions was challenging, as well as locating genes within a bacterial genome. More recently, tools have also been developed to allow these genomes to be constructed, and these are referred to as metagenomically-assembled genomes (MAGs). This represents a further major step change in our understanding of microbiomes. In the context of the rumen microbiome, there are now 4,941 MAGs available, representing a major resource for enhanced understanding and for compound/enzyme mining purposes.
Agricultural microbiomes: what does the future entail?
Of course, while the field has progressed substantially, it is important that we don’t ‘oversell the microbiome’, a concept used by Dr Jonathan Eisen (University of California, Davis), in which he awarded prizes on social media for publications that did exactly that. This is largely a response to correlative studies and wild statements resulting from those publications without definitive proof. Certainly, in the agricultural setting, there is a need for a major push towards causative, rather than correlative studies, as few studies are available that can actually show causality. Developments in MAG isolation also provide a mechanism to move towards causality, as they provide information on the likely biochemical needs of a bacterium, allowing more informed culture-based technologies to be developed. Ultimately, having availability of pure cultures is crucial to develop hypotheses to confirm causality and the recent surge in culturomic-based activity is highly commended. In conclusion, the explosion in ‘omic technologies and datasets available for agriculture-based microbiomes has been somewhat overwhelming in recent years. The lack of standardisation or controls used in studies makes comparisons between studies difficult and ultimately decreases our ability to look at global datasets using meta-analyses to increase the power of observations across geographical locations. While standardisation is challenging between laboratories in different geographical locations, the use of controls is critical for the future as we move from correlative studies to defining causality.

Sharon Huws
School of Biological Sciences/Institute of Global Food Security, Queen’s University Belfast, 19 Chlorine Gardens, Belfast BT9 5DL, UK
[email protected]
+44 (0)28 9097 2412
@SharonHuws
Sharon Huws is a Professor in Animal Science and Microbiology at Queen’s University Belfast, UK. Professor Huws specialises in understanding livestock-related microbiomes in relation to their influence on animal production, health and the environment. She is also Editor-in-Chief for the journal Animal Microbiome, a sister journal to Microbiome.
Why does microbiology matter?
Most metabolic processes in nature are underpinned by microbial activity and industrial processes are enabled by the actions of microbes and their enzymes. Therefore, microbiology has always been and always will be at the core of most scientific endeavour, with few disciplines having such vast interdisciplinary importance.
What advice would you give to someone starting out in this field?
I would say don’t worry if you take some jagged routes to your final destiny. My path into microbiology has been a somewhat jagged one, and this jagged route is not one I regret as it’s helped to develop systems-level thinking beyond the discipline. I’d also say take time to think – while it may seem an impossible thing to do in the ‘race’ towards qualifications, publications and obtaining a permanent position early in your career. Obviously, this is a utopian ideal and many scientists have other responsibilities, such as having a family, but it gets much more challenging as your career develops, therefore, make the most of this time.
Images: Guardians of the Gut’. Lindsay Hall.
Louis Pasteur, famous for the discovery of yeasts as agents of fermentation.
Illustration from 1894 stock illustration. suteishi/iStock.
Fig. 1. Sharon Huws.
Find out more about 'Unlocking the world of microbiomes: exploring microbial communities' in our digital content hub.
-
Unlocking the world of microbiomes: exploring microbial communities
Research into the microbiome has evolved over time allowing us to study microbial communities, genes and proteins in more depth. We will explore three key areas of microbiome research: the microbiome and human health, agricultural and food microbiomes and environmental and industrial microbiomes.
Image: iStock/Martina Birnbaum.