Drugs from bugs that kill bugs
Issue: Antimicrobials
01 November 2012 article
HELGE B. BODE
It isn't just soil-dwelling actinomycetes that may hold the key to new antimicrobial compounds. Insect-pathogenic bacteria also produce a host of bioactive compounds – and what's more, they are easy to analyse in vivo.
Natural products from microbes and plants have been used in human medicine for thousands of years. During the last 60 years they have been especially useful as antibiotics and anti-cancer compounds. Most low-molecular-mass compounds used clinically are in fact natural products, natural product derivatives or have been inspired by natural products. However, the current situation where we have antibiotics to treat many bacterial infections is likely to change rapidly in the next decade or two due to increasing resistance and the emergence of not only multi-, but pan-resistant human pathogens. At the same time, pharmaceutical companies have stepped back from antibiotic research due to high development costs and have concentrated mostly on drugs for chronic diseases.
This dramatic situation in the clinic contrasts with the enormous possibilities of modern science. Next-generation sequencing allows cheap and rapid access to biosynthesis gene clusters in bacteria and fungi, which can be manipulated very efficiently in the original host using genetic tools, or expressed heterologously in more accessible model organisms, such as Escherichia coli, Bacillus or optimised Streptomyces hosts. Additionally, analytical methods, like mass spectrometry, allow the rapid detection and structural elucidation of novel natural products or their derivatives. The development of these tools has already led to the revitalisation of academic natural product research.
PHOTORHABDUS AND XENORHABDUS ARE POTENT NATURAL-PRODUCT-PRODUCERS
Detailed analysis of well-known antibiotic producers like Streptomyces has shown that their potential to produce novel compounds is still enormous. In addition, it has been shown over the last few years that several other bacterial phyla are in fact promising natural-product-producers.
Five years ago, we started to analyse bacteria of the genera Photorhabdus and Xenorhabdus for their ability to produce natural products, as earlier work had revealed the presence of compound classes unique to these bacteria.
Photorhabdus and Xenorhabdus belong to the Enterobacteriaceae and live in symbiosis with nematodes of the genera Heterorhabditis and Steinernema, respectively. Both bacterium and nematode form an entomopathogenic complex that can infect and kill several soil-dwelling insect larvae and therefore this complex can be used industrially for pest control
in organic farming. The nematode can be regarded as the carrier for the bacteria, setting them free once inside the insect, and the bacteria essentially function as the warhead that kills the insects.
Several protein toxins are major players in carrying out their deadly job; there are also a number of low-molecular-mass natural products with insecticidal activity that have also been isolated from these bacteria.
Another peculiarity of Photorhabdus is its bioluminescence,
which makes it unique among terrestrial bacteria, and it is already an established tool in modern molecular biology, although its true biological function is still not understood.
During the course of the EC-funded project ‘Genomic approaches to metabolite exploitation from Xenorhabdus/Photorhabdus’ (GameXP) with partners from the UK (Sharon Peacock, Cambridge; Nick Waterfield, Bath; Richard ffrench-Constant, Exeter), Thailand (Narisara Chantratita, Bangkok), Vietnam (Long Phan Ke, Hanoi) and Germany (Vera Siegmund, Saarbrücken), the chemical diversity of these bacteria was investigated and exploited.
More than 200 different bacterial strains were isolated from nematode-infected insects obtained from soil samples (some together with their associated nematodes) and the bacterial strains were analysed using mass spectrometry. Using bioinformatics, over 500 novel compounds have been identified in these bacteria and more than 150 compounds have been isolated and/or synthesised.
These compounds range from simple amino acid derivatives like phenylalanine-derived cinnamic acid or simple amides to very large peptides with a molecular mass greater than 1,800Da. All compounds have been tested against different insects or insect cells, pathogenic bacteria and parasites such as Leishmania (leishmaniasis), Trypanosoma (sleeping sickness and Chagas disease) or Plasmodium (malaria).

Different insecticidal and antibiotic compounds have been identified and, more surprisingly, a high proportion of compounds showed activity against the human parasites tested, and some of these are being analysed in detail using in vivo models.
Another goal of GameXP was to improve the production of these bioactive compounds and to increase the chemical diversity. As the regulation of the promoters that drive the expression of these compounds is often not understood, these promoters have been exchanged with strong constitutive or inducible promoters, enabling ‘on demand’ and high-titre production of selected compounds. With the genomes of several chemically diverse strains in hand and several biosynthesis gene clusters responsible for the production of typical natural products like peptides and polyketides already identified, the promoter exchange approach can now be applied to several strains.

The high degree of bioactive compounds in general may be the result of the specific ecological niche these bacteria inhabit. Being nematode symbionts they must be able to switch between a mutualistic (towards the nematode host) and a pathogenic (towards the insect prey) growth phase; this may require the production of specific small-molecule effectors, signals or toxins. Additionally, they have to protect the dead insect from food competitors, such as soil-dwelling bacteria, fungi and amoebae. Thus, the biochemical similarity between amoeba and human parasites, such as Plasmodium, may be the reason for the bioactivities observed.
One major question is how do the bacteria know where they are and what compounds are to be produced in their given environment? The group of Jon Clardy in Harvard has shown that insect blood might trigger the production of some (insecticidal?) natural products. Further triggers are likely to be discovered in the future as these bacteria contain a variety of different sensor systems that might be involved in integrating different environmental cues.
FINDING THE TRUE FUNCTION OF NATURAL PRODUCTS
Another advantage of these bacteria is that the real function of natural products can be studied. Although natural products are indeed important therapeutics, the function and regulation mechanisms leading to their production are mostly unknown. It is clear that bacteria in general do not produce these compounds for the purpose of selling them in a pharmacy! To analyse the natural function of compounds in established producers such as Streptomyces or Bacillus is more challenging than in Photorhabdus or Xenorhabdus, as we know only a fraction of their possible interaction partners in their natural environment, the soil. Photorhabdus and Xenorhabdus can be grown in the lab alone or together with their interaction partners (nematodes, insects or food competitors), allowing the analysis of all possible growth phases of these bacteria using analytical and molecular biology methods. It is evident that some of these natural products must play an important role in mediating these complex interactions from their conservation in several distantly related bacterial strains.
In summary, GameXP has placed Xenorhabdus and Photorhabdus in the company of other well-known natural-product-producers like Streptomyces, Bacillus and myxobacteria. Although their compounds still have much to prove in terms of potency as clinical antibiotics or anti-cancer compounds, their potential for therapeutic use is very high.
ACKNOWLEDGEMENTS
The author is grateful to the GameXP team and all members of his group for their enthusiasm and fun during recent years. The work was supported by the EC’s Seventh Framework Programme (FP7/2007-2013) under grant agreement no. 223328, the German Research Council (DFG) and the LOEWE excellence program Insect Biotechnology.
HELGE B. BODE
Merck Stiftungsprofessur für Molekulare Biotechnologie, Fachbereich Biowissenschaften, Goethe Universität Frankfurt Max-von-Laue-Str. 9, 60438 Frankfurt am Main, Germany;
Tel. +49 69 798 29557;
Email [email protected]
FURTHER READING
Bode, H.B. (2009). Entomopathogenic bacteria as source of new secondary metabolites. Curr Opin Chem Biol 13, 224–230.
Crawford, J.M. & Clardy, J. (2011). Bacterial symbionts and natural products. Chem Comm 47, 7559–7566.
Chaston, J.M., Suen, G., Tucker, S.L. & others (2011). The entomopathogenic bacterial endosymbionts Xenorhabdus and Photorhabdus: convergent lifestyles from divergent genomes. PLoS One 2011 6, e27909.
Reimer, D., Pos, K.M., Thines, M., Grün, P. & Bode, H.B. (2011). A natural prodrug activation mechanism in nonribosomal peptide synthesis. Nat Chem Biol 7, 888–890.
FIGURES
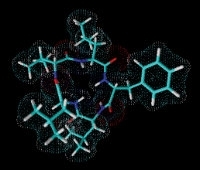
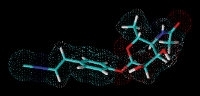
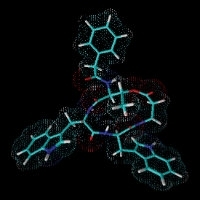
GameXPeptide A, a cyclic pentapeptide from Photorhabdus
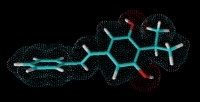
Isopropylstilbene from Photorhabdus required for Heterorhabditis development. Photorhabdus is the only non-plant organism able to produce stilbenes.
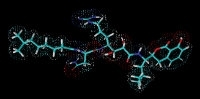
Prexenocoumacin C from Xenorhabdus nematophila. Prexenocoumacins are inactive prodrugs of the potent antibiotic xenocoumacin-1, which result from cleavage by a specific peptidase.
Rhabduscin produced by Xenorhabdus and Photorhabdus that inhibits phenoloxidase, which is part of the insect immune system.
Xenematide from Xenorhabdus nematophila, an insecticidal depsipeptide. H.B. Bode
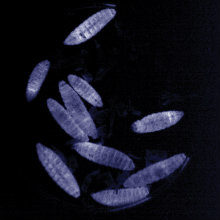
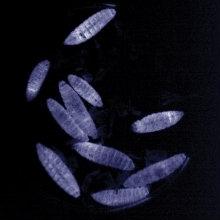
Larvae of the Greater waxmoth (Galleria mellonella) infected with Photorhabdus showing the typical bioluminescence. G. mellonella is used as a model for bacterial infections. H.B. Bode
Tobacco hornworm (Manduca sexta) caterpillar used as a model for bacterial infections. H.B. Bode
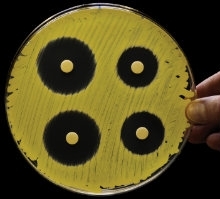
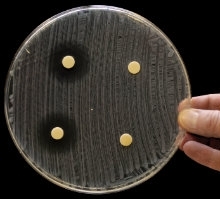
Inhibition zones of compounds from Xenorhabdus against Micrococcus luteus. H.B. Bode
Inhibition zones of compounds from Xenorhabdus against E. coli. H.B. Bode
