A plague on our ashes
Issue: Fungal diseases
09 February 2016 article
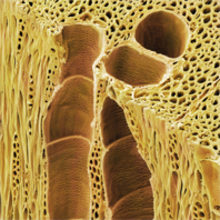
In late autumn 2012, it became evident that ash dieback disease had become established in native woodlands in the eastern regions of the UK. This was the leading edge of an epidemic that had already swept east to west across mainland Europe over a period of about 20 years.
We were aware of the disease because of our involvement in helping manage Ashwellthorpe Lower Wood, an ancient coppiced woodland in South Norfolk. This woodland, owned by the Norfolk Wildlife Trust, is a Site of Special Scientific Interest (SSSI) and its name, Ashwellthorpe, comes from the Vikings, for whom the ash (Fraxinus excelsior) was sacred. It was devastating to see young coppiced regrowth showing early signs of what looked like ash dieback. Since this occurred just a few miles from where we work at the John Innes Centre (which specialises in plant and microbial research), we asked ourselves and our colleagues what we could do to help an iconic native tree.
As a first step, we extracted and tested DNA from infected branches and our fears were confirmed: the trees were infected with the fungus Chalara fraxinea, the causal agent of ash dieback. Symptoms of the disease include the wilting (dieback) of young leaves in spring, dying of leaves in late summer while they are still attached to the tree, lesions and cankers on stems and branches, and dieback of the crown. Young saplings and coppice regrowth can succumb quite rapidly, some within one growing season. Older trees can survive an initial attack but many stop growing and may become prone to secondary infections. In other parts of Europe, over 90% of the ash trees were killed or severely affected by the disease, so there was an outcry in the UK as the general public and media realised that this iconic tree may disappear from our landscape.
WE REVISIT THE WOODLAND WHERE ANNE FIRST FOUND ASH DIEBACK IN 2012, TO SEE HOW SOME TREES SEEM TO BE TOLERANT TO C. FRAXINEA, AND LEARN WHAT'S BEING DONE IN THE FIGHT AGAINST THE FUNGAL PATHOGEN.
A new pathogen in Europe
What was already known about the disease-causing fungus? Scientists from across Europe had established that Chalara fraxinea is the vegetative (anamorphic) form of the fungus, which kills trees by invading stems and trunks and blocking the vasculature. The fungus also has a sexual form, which was originally named Hymenoscyphus pseudoalbidus, because it looked similar to, and has the same host as, Hymenoscyphus albidus, which grows widely (but non-pathogenically) on ash trees across Europe. Due to its distinctive characteristics, the pathogen was renamed Hymenoscyphus fraxineus, and we now know that where the disease is present, H. fraxineus is completely displacing H. albidus. Both species live in ash leaf litter and spread by releasing sexual (haploid) spores into the air and infecting surrounding trees. Spores fired upwards from fruiting bodies land on ash leaves, germinate and penetrate as the vegetative form into the petiole and main leaf stem (rachis). When the rachis falls, it takes the fungi with it and sometime during the winter and following spring, the fungi enter the sexual phase of the lifecycle. Pre-fruiting structures start to appear, some of which develop into tiny white cup-shaped mushrooms (ascocarps) less than 0.5 cm wide. There can be 20 of these mushrooms on a single petiole and when they mature in summer, they release highly infective ascospores (around 1,500 per hour every morning in summer months) and the whole cycle starts again.
Normally, trees rid themselves of the saprophyte H. albidus in autumn with leaf fall. However, H. fraxineus grows faster and more aggressively on ash trees and so, rather than being contained within the leaves and leaf stems, H. fraxineus can grow into the woody parts of the tree. There it continues to grow slowly over winter and then in spring probably grows rapidly as the tree starts to initiate new leaf growth. This rapid fungal growth in the phloem and xylem of branches can then strangle the growth of the distal young leaves, causing the characteristic ‘dieback’ symptoms.
Had the harmless native saprophyte turned into a killer or was the destructive H. fraxineus itself a saprophyte which had found itself in a foreign environment without its natural host? The latter theory gained popularity when it was found that Asian ash species such as Fraxinus mandschurica were unaffected by ash dieback. Genome sequencing has revealed that H. fraxineus has a considerably larger genome (60 megabases) than the saprophyte H. albidus (40 megabases) and so these are two related but rather different species. It is probable that H. fraxineus was introduced into Eastern Europe, possibly due to movement of trees or leaves from somewhere in East Asia, where F. mandschurica is adapted to growth with H. fraxineus.
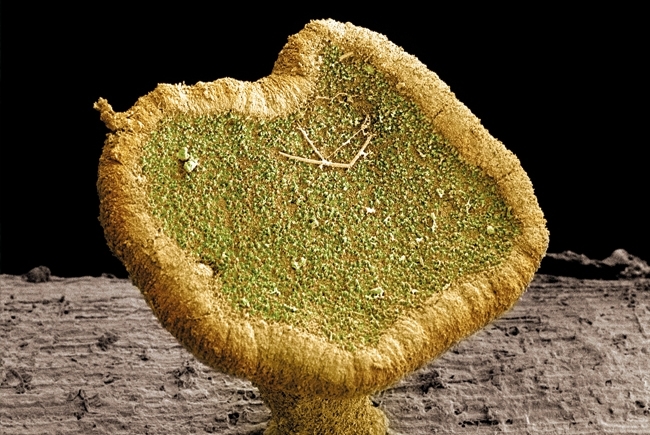
Genetic tolerance in ash trees
F. excelsior can respond in different ways when challenged by H. fraxineus. Many trees appear to be exceptionally sensitive, dying within a few years of infection; a few others (about 5% in Denmark) seem to be much more tolerant to the disease. In between these two extremes, there is a range of responses. Danish scientists had grown several clones of grafted trees in many different environments and fortuitously a couple of these showed some tolerance (one high and one intermediate) to the disease. Since this tolerance was consistently observed in various locations and in different environments, it could be concluded that the tolerance was genetically determined.
Genomics of host and pathogen
With funding from the BBSRC and Defra, a research consortium called Nornex was set up including groups from the John Innes Centre, The Sainsbury Laboratory and The Genome Analysis Centre (all in Norwich), the Universities of Exeter, Edinburgh and York, FERA, Forest Research, the University of Copenhagen, and the Norwegian Forest and Landscape Institute. The aims were to (a) use genomics-based tools to establish a foundation of knowledge on the pathogen, (b) establish laboratory-based tests of pathogenicity and assays of growth and survival of the pathogen, and (c) use genomics and transcriptome-based approaches in ash trees to try to map tolerance (resistance) to the disease.
These aims were underpinned by a desire to try to take a different approach and use an open access and crowdsourcing approach. As part of that, a website called openashdieback was established to host new observations and provide access to data generated by the consortium. In view of the intense public interest, we also developed and released Fraxinus, a Facebook-based citizen science genomics-based game. This enabled non-specialists to contribute to the genomic studies of the pathogen by helping to improve genetic variant predictions from DNA sequence alignments. Many computer programs have been devised to optimise alignments. However, none can improve on the pattern recognition skills of the human eye. Once an alignment process is complete, genetic variations such as single nucleotide polymorphisms (SNPs) or insertion-deletion polymorphisms (INDELs) can be identified and further studied in relation to alterations in, for example, pathogenicity.

The research
Three years on and what has been achieved? We have a good genome sequence of H. fraxineus, revealing that it does not contain the great diversity of genes for wood degradation like those used by wood-rotting fungi, but it does possess cell wall-degrading enzymes such as cellulases and pectate lyases used for cellular penetration. It also synthesises toxins, effectors and predicted protein inhibitors to help it evade plant defences. Genome sequencing of different isolates has revealed that there is less genetic diversity in European isolates than in Asian isolates and it seems highly likely that the original H. fraxineus came from Asia. We have preliminary data from the sequencing of tree leaf RNA, identifying potential genetic markers that could be associated with inheritance of tolerance to the disease. We believe that we have established a framework that will enable us to predict which trees will be tolerant to the disease, to understand the nature and diversity of the pathogen, and to carry out laboratory-based tests of pathogenicity in tree seedlings. All of these can be used to rebuild a genetically diverse population of trees tolerant to this disease that, over time, will probably kill or severely damage 80–90% of the UK ash tree population.
ANNE EDWARDS AND J. ALLAN DOWNIE
John Innes Centre, Norwich Research Park, Norwich NR4 7UH, UK
[email protected]
[email protected]
FURTHER READING
Gross, A. & others (2012). Reproductive mode and life cycle of the ash dieback pathogen Hymenoscyphus pseudoalbidus. Fungal Genetics and Biology 49, 977–986.
McKinney, L. V. & others (2011). Presence of natural genetic resistance in Fraxinus excelsior (Oleraceae) to Chalara fraxinea (Ascomycota): an emerging infectious disease. Heredity 106, 788–797.
Saunders, D. & others (2014). Crowdsourced analysis of ash and ash dieback through the Open Ash Dieback project: A year 1 report on datasets and analyses contributed by a self-organising community. BioRxiv. doi:10.1101/004564.
Rallapalli, G. & others (2015). Cutting edge: Lessons from Fraxinus, a crowd-sourced citizen science game in genomics. eLIFE. doi:10.7554/Elife.07460.
Image: Scanning electron micrograph of the wood of an ash tree (Fraxinus excelsior) showing xylem tracheids. Power and Syred/Science Photo Library. Coloured scanning electron micrograph of a fruiting body of the fungus Chalara fraxinea (Hymenoscyphus pseudoalbidus), the causal agent of ash dieback. Crown © Courtesy of FERA/Science Photo Library. Fraxinus, a Facebook-based citizen science genomics-based game. Anne Edwards and Allan Downie..
