Going green through co-operation: the origins of chloroplasts
Issue: Light
11 August 2015 article
‘Life did not take over the globe by combat, but by networking’
Lynn Margulis
Machines of life: from cyanobacteria to chloroplast
Light. It’s all around us. We rely on plants and algae to make use of its energy, through the process of photosynthesis. Photosynthesis within bacteria evolved early in Earth’s history. Around 3.6 billion years ago, cyanobacteria (blue-green algae) adopted the main form of photosynthesis we see today, in which carbon dioxide and water are converted, using captured sunlight, into energy-rich sugars and oxygen.
This major biological feat transformed the geochemistry of the planet, enriching the atmosphere with oxygen, and shaping the biosphere as we know it today. At the same time, photosynthetic organisms have diversified. Not only is photosynthesis found within the bacteria, it is used by a wide range of eukaryotes, from unicellular algae and giant seaweeds in the oceans to plants that flourish on land. What is common to all these eukaryotes? Cyanobacteria-like chloroplasts.
The merge: when two became one
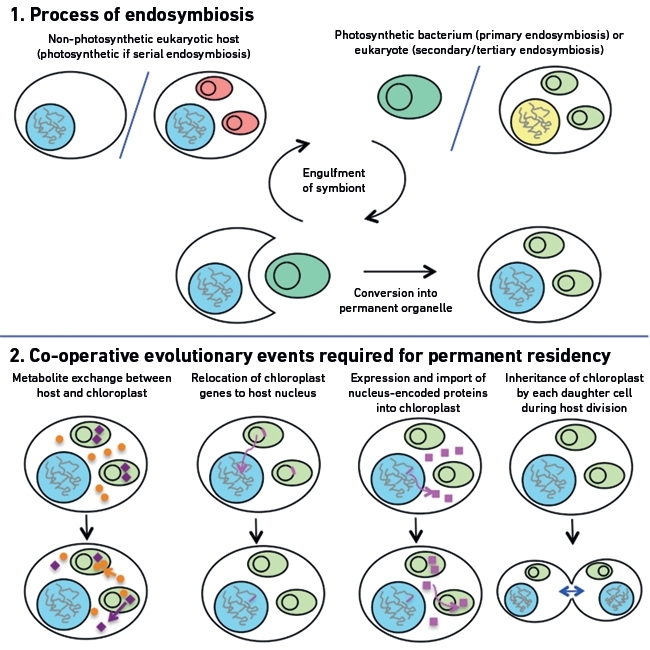
Microscopists have long recognised that the structure and staining properties of chloroplasts are very similar to those of some free-living bacteria. In 1910, the Russian biologist Mereschkowsky postulated that chloroplasts not only resembled prokaryotes, but were indeed remnants of once free-living bacteria. Mereschkowsky’s ideas were developed further in the 1960s by Lynn Margulis. Margulis theorised that an early eukaryote engulfed a free-living cyanobacterium, and converted it into a cellular organelle (Fig. 1). This process is called endosymbiosis.
Margulis’ hypothesis was initially viewed with much scepticism. Before her ideas were eventually published in 1967, they had been rejected by fifteen scientific journals! However, experimental evidence has since left the theory uncontested. Most significantly, chloroplasts possess their own genomes, and these have been shown to be closely related to the genomes of cyanobacteria. Today, it is widely accepted that chloroplasts evolved through the merging of two cells into a single lineage.
The permanent house-guest: integration and streamlining
Since the initial endosymbiotic event, the host cell and chloroplasts have had to learn to co-operate with one another, setting the stage for a permanent residency (Fig. 1). Chloroplasts, for example, supply the host with sugars and other compounds synthesised through photosynthesis, and the host supplies essential metabolites and co-factors to the chloroplast (Fig. 1). In addition, the host physically controls the chloroplast, coordinating its replication, and protecting it from environmental stresses, ensuring that each daughter cell inherits a functional chloroplast (Fig. 1).
This physiological integration has been underpinned by changes to the genomes of the host nucleus and the chloroplast. Most dramatically, chloroplast genomes have undergone massive streamlining. While free-living cyanobacteria contain upwards of 3,000 genes, chloroplast genomes generally retain fewer than 250.
Many of the genes that were streamlined from chloroplast genomes have been transferred into the nucleus of the host (Fig. 1). Remarkably, the host cell was able to re-introduce the expression products of some of these genes, now translated in the cytoplasm, back into the chloroplast (Fig. 1). Nucleus-encoded chloroplast proteins gain targeting sequences that guide the protein ‘home’, across a complex protein import machinery. These nucleus-encoded, chloroplast-originated proteins allow the host to directly control the physiology of the chloroplast through altering nuclear gene expression. In addition, the host has adapted some of its own genes to also encode chloroplast-targeted proteins. This has allowed the host to effectively ‘customise’ the biology of the chloroplast to suit its needs.
FIG. 1. THE ENDOSYMBIOTIC EVOLUTION OF CHLOROPLASTS FROM CAPTURED MICROBES.
A rich tapestry of evolution: diversity and promiscuity
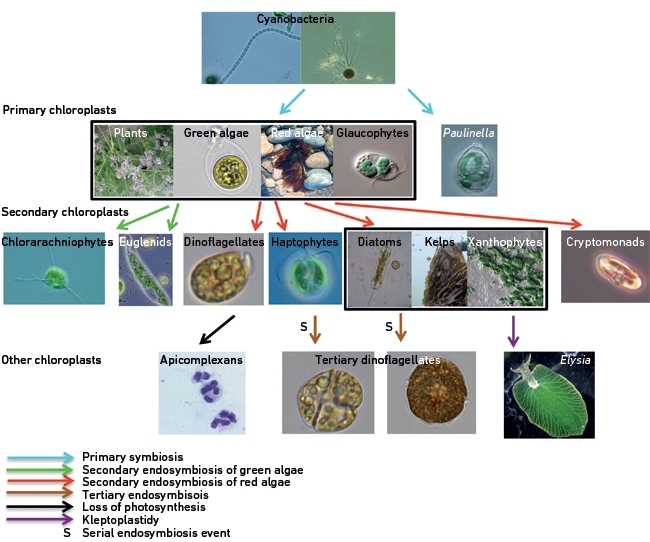
The endosymbiotic integration of cyanobacteria has spawned an overwhelming array of chloroplast lineages. A single primary endosymbiosis gave rise to three groups: the green algae (and their descendants, the plants), the red algae (which includes the edible seaweed nori), and the glaucophytes (Fig. 2).
However, the majority of algal diversity comes from another form of genomic gymnastics: secondary endosymbiosis. Here, a new host engulfed an existing red or green alga, already containing a primary chloroplast, to form a eukaryote–eukaryote partnership, effectively a ‘meta-alga’. The ‘meta-algae’ include environmentally important lineages, such as kelps, diatoms, haptophytes (who also form the principal component of chalk) and dinoflagellates (which, amongst other roles, are the photosynthetic component of corals) (Fig. 2). Perhaps most atypical within the ‘meta-algae’ is the malaria pathogen Plasmodium, a member of the apicomplexan group (Fig. 2). This group of eukaryotes descended from algae but has secondarily lost the ability to photosynthesise, and instead survives through parasitism (Fig. 2).
Even more unusual chloroplasts are found elsewhere. Some dinoflagellates have chloroplasts that appear to be derived through the ingestion of an alga with a secondary chloroplast (diatoms, and haptophytes), in a proposed ‘tertiary endosymbiosis’ (Fig. 2). These chloroplasts are also believed to have arisen through ‘serial endosymbiotic’ events, in which a dinoflagellate that possessed secondary chloroplasts underwent a subsequent endosymbiotic event, swapping its original chloroplasts with new chloroplasts of a novel phylogenetic origin. It is now debated whether some other chloroplast lineages, such as those of diatoms, might have also actually arisen through complex tertiary and serial endosymbiotic events.
Weird and wonderful more recent symbioses: kleptomaniacs
The primary chloroplast endosymbiosis and their secondary endosymbiotic derivatives have had profound effects on eukaryotic life. However, in addition to these conventional chloroplasts, a number of organisms demonstrate that endosymbiosis is a continual driving force in evolution, and give us insights into how chloroplasts first evolved.
Paulinella chromatophora is a freshwater amoeba, which contains chloroplasts (Fig. 2). Surprisingly, these chloroplasts are unrelated to all other chloroplast lineages and instead appear to have originated from a separate primary endosymbiosis, involving a different cyanobacterial symbiont. Notably, examples have been documented in Paulinella of genome streamlining, gene transfer to the nucleus and the import of nucleus-encoded proteins into the chloroplast, cementing their importance for the conversion of chloroplasts into permanent organelles.
Perhaps the most weird and wonderful chloroplast lineage are the stolen chloroplasts (‘kleptoplasts’) found in the green sea slug Elysia chlorotica. Young Elysia feed on the alga Vaucheria litorea, and harvest the chloroplasts. These are maintained over the entire lifespan of the sea slug, which needs not feed again (Fig. 2). While it is not certain whether these chloroplasts are functional inside the host, or are merely stored as food resources, a recent fluorescent labelling study has indicated the presence of an algal gene on an Elysia chromosome, suggesting gene transfer has happened again. Could this intricate partnership represent an ‘endosymbiosis in progress’? Only (a lot of) time will tell!
FIG. 2. THE DIVERSITY OF PHOTOSYNTHETIC EUKARYOTES. EACH ARROW CORRESPONDS TO A SEPARATE PROPOSED ENDOSYMBIOTIC EVENT. LINEAGES THAT ARE SHOWN SURROUNDED BY A BLACK BOX ARE BELIEVED TO HAVE ARISEN THROUGH THE SAME ENDOSYMBIOSIS. IMAGES ARE NOT TO SCALE.
Conclusion
A traditional view of evolution is one driven solely by competition and a vertical transmission of genes. The fascinating world of endosymbiosis throws a spanner in the works, shedding light on complex partnerships, which blur the boundaries between organisms. The capture of a photosynthetic cyanobacteria-like prokaryote over a billion years ago paved the path for eukaryotes to thrive on the Sun’s energy, and transformed life as we see it today.
PHOEBE TICKELL
Department of Life Sciences, Imperial College London, South Kensington Campus, London SW7 2AZ, UK
RICHARD G. DORRELL
Department of Biology, École Normale Supérieure, 46, Rue d’Ulm, 75005 Paris, France
[email protected]
ACKNOWLEDGMENTS
The authors would like to thank Dr Leila Tirichine and Dr Zhanru Shao (École Normale Supérieure); Professor Alison Smith and Dr Katherine Helliwell (University of Cambridge); Dr Nic Blouin (University of Rhode Island); and the contributors to Micro*scope (http://pinkava.asu.edu/starcentral/microscope) and Encyclopedia of Life (www.eol.org), for the kind provision of images featured in Fig. 2.
FURTHER READING
Dorrell, R. G. & Smith, A. G. (2011). Do red and green make brown? Perspectives on plastid acquisitions within chromalveolates. Euk Cell 10, 856–868.
Nakayama, T. & Archibald, J. M. (2012). Evolving a photosynthetic organelle. BMC Biol 10, 35. doi:10.1186/1741-7007-10-35.
Schwartz, J. A., Curtis, N. E. & Pierce, S. K. (2014). FISH labelling reveals a horizontally transferred algal (Vaucheria litorea) nuclear gene on a sea slug (Elysia chlorotica) chromosome. Biol Bull 227, 300–312.
Image: Light micrograph of Glaucocystis sp. algae. Michael Abbey/Science Photo Library. Fig. 1. R. Dorell and P. Tickell. Fig. 2. R. Dorell/P. Tickle/Micro*scope/Encyclopedeia of Life..
