A special sugar: how sialic acid impacts on metabolism, health and disease
Issue: Metabolism, Health and Disease
07 May 2019 article
We are teeming with microbes that live on surfaces outside and inside our bodies. Our understanding of the gut microbiome in particular is rapidly improving as we start to unpick how complex microbial communities interact with our diet, with other microbes and with our cell surfaces. While many chemicals are important in this milieu, here we focus on sialic acids as a critical set of molecules that underpin many of these interactions and impact on metabolism, health and disease.
The most common sialic acid, N-acetyl-neuraminic acid (Neu5Ac) (Fig. 1) is a relatively simple sugar acid that, apart from its charge, has nothing intrinsically unusual about it. However, when cells contact other cells they often use molecules displayed on the surface, and this is where sialic acids become so important, as they are usually the terminal sugars on the host glycans that pepper proteins and lipids sitting on cell surfaces. Cell-surface or secreted mucin glycoproteins are good examples of sialic acid-coated structures that microbes could encounter at mucosal surfaces, such as the respiratory or gastrointestinal (GI) tract. This unique position as the ‘meet and greet’ molecule for other human cells, bacterial cells and viruses puts the sialic acids at centre-stage for many important processes.
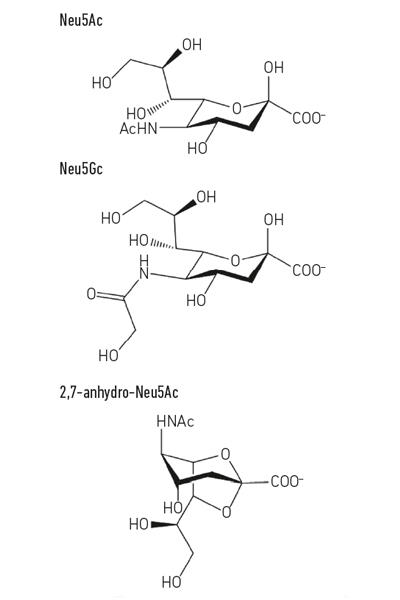
Fig. 1. Example of sialic acid derivatives. Note that the N-glycolyl-neurnaminic acid (Neu5Gc) cannot be made by humans, a biochemical difference that distinguishes us from other great apes, but can be incorporated into our cell surfaces when acquired from the diet. |
|
Sugar-coated pathogens
One of the first microbial functions of sialic acids was connected to the ability of some pathogens to colonise and then cause disease in humans. Indeed, the ex-President of the Microbiology Society, Professor Harry Smith FRS, working at the University of Birmingham, pioneered this work with Neisseria gonorrhoea. When scientists in his group isolated the bug directly from the body, it was resistant to killing by human serum (complement-mediated killing), but if the bugs were cultured in the lab for a few generations, this protection was lost and the cells were killed by the serum. His group had discovered that these pathogens steal sialic acid from the host and use it to coat their own cell surface to humanise them. However, as they can only scavenge this sialic acid, its absence in growth media means no further sialic acid is added so they rapidly lose this ‘serum protection’ and are killed. It turns out that a range of other sneaky bugs use similar strategies to generate their own invisibility cloak using sialic acid, such as Neisseria meningitidis, Campylobacter jejuni or Haemophilus influenzae (Fig. 2a)
Cross-feeding – sharing the sweets around
As well as being used by pathogens as an invisibility cloak, sialic acids are pretty good food for bacteria, as they provide both carbon and nitrogen, and can be used as an energy source. Earlier work showed that pathogens such as Vibrio cholerae were happy just to eat sialic acids and that in itself was important for successful host colonisation (Fig. 2b).
In the gut, a major source of sialic acids comes from mucins, which are the main structural components of the mucus layer covering the epithelium surface. The most exposed layer of mucus is the habitat of various commensal bacteria, which have learnt how to feast on mucin proteins, where sugars make 80% of their mass. From a simple topological perspective, the sialic acids, as the terminal sugars, are harvested first, exposing the underlying sugars that can also be subsequently released and eaten. With the advent of next-generation sequencing, we have gained knowledge on the microbes living in this niche, and many have genes for sialic acid uptake and subsequent catabolism. What was exciting was the discovery that some of the commensal anaerobes, like Bacteroides thetaiotaomicron, secrete sialidases to release the sialic acid but lack transporters and catabolic genes to actually use it. This seemingly altruistic gesture is required for the bacteria to access underlying sugars and underpins a process of metabolic cross-feeding to other bugs living in the mucus niche, like commensal Escherichia coli strains that do not have a sialidase but are good at eating free sialic acid (Fig. 2c). This concept is now recognised as an important aspect of colonisation and niche adaptation. A brilliant exemplification of this was described by the group of Justin Sonnenburg from Stanford University, using a mouse model with a simple defined gut microbiota. Following antibiotic treatment, many of the sialic-acid-eating bacteria were wiped out, resulting in the accumulation of free sialic acid in the gut. When then challenged with pathogens like Salmonella typhimurium or Clostridium difficile, these pathogens would benefit from the free sialic acid as a nutrient (Fig. 2d). Mutant strains of pathogens lacking the transporters or catabolic genes did not have an advantage, making a direct connection to pathogen outgrowth when the gut microbial community is disrupted. Mechanisms to reduce the pool of free sialic acid in the gut were therefore proposed as part of a novel anti- infective strategy.
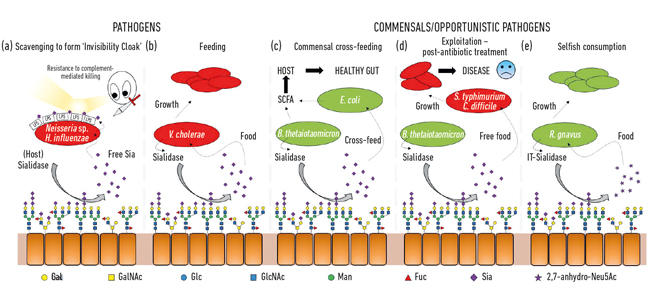
Fig. 2. Cartoon illustration of the various interactions between bacteria and sialic acids present as terminal sugars on a range of host glycans. These are cleaved by sialidases and usually converted to free Neu5Ac with the exception of the IT-sialidase that releases 2,7-anhydro-Neu5Ac. Free sialic acid released by host or pathogenic sialidases can then be used by pathogens as an invisibility cloak (a), or for feeding (b). Cross feeding can occur between members of the gut microbiota or between commensal and pathogenic bacteria sharing the same niche (c, d) while other bacteria have developed selfish mechanisms to reserve sialic acid for their own consumption (e).
Keeping hold of your own sweets
While much early sialomicrobiology focused on pathogens, more recent work has focused on the commensal bacteria inhabiting the same niche. Today’s studies in the field of the gut microbiota are revealing how fierce the competition for Neu5Ac is in our gut and how it can tip the balance between pathogens and commensals. What if a particular bacterium could release sialic acid in a form that only it could access? Work in the Juge Group in Norwich showed that the gut symbiont Ruminococcus gnavus has evolved to do so, using a special type of sialidase called an intramolecular trans-sialidase (IT-sialidase) which releases 2,7-anhydro-Neu5Ac (see Fig. 1) instead of Neu5Ac. By developing a method to synthesise pure 2,7-anhydro-Neu5Ac, they showed that R. gnavus strains were able to grow on 2,7-anhydro-Neu5Ac as the sole carbon source. In addition to the domain catalysing the enzymatic reaction, IT-sialidases comprise a carbohydrate binding domain which helps to mediate the binding of the IT-sialidase to sialic acid rich mucins. In vivo, this may favour a mechanism that helps targeting the bacteria towards sialic acid rich regions of the GI tract, therefore promoting bacterial colonisation within the mucus layer. Once located in the right place, the IT-sialidase cleaves off sialic acid from the terminal chains of mucins and releases it in a form, 2,7-anhydro-Neu5Ac, that R. gnavus can use as its own source of food, rather than sharing Neu5Ac with other bacteria inhabiting the mucus niche, including pathogens (Fig. 2e). This mechanism provides a competitive advantage over other sialic-acid- eating bacteria inhabiting the mucus niche, by providing a source of nutrient that it can preferentially access. In addition, R. gnavus may also help reduce outgrowth of enteric pathogens by reducing the level of Neu5Ac in the mucosal environment, opening ways to a novel therapeutic strategy in an age of increasing antibiotic resistance.
Viruses get in on the act |
|
|
In conclusion, it is clear that sialic acids are critical molecules in many processes where microbial cells and viruses interact with mammalian cell surfaces. In this short article, we have touched on some of these to give an idea of the central functions of these sugars in communication, stealth, colonisation and metabolism in different aspects of heath and disease. Sialic acids have been described as the most important chemicals in the world and will no doubt continue to reveal more about the evolution of humans and their interplay with their microbes.
Further reading
Thomas, GH. Sialic acid acquisition in bacteria – one substrate, many transporters. Biochem Soc Trans 2016;44:760–765.
Ng KM, Ferreyra JA, Higginbottom SK, Lynch JB, Kashyap PC et al. Microbiota-liberated host sugars facilitate post-antibiotic expansion of enteric pathogens. Nature 2013;502:96–99.
Owen CD, Tailford LE, Monaco S, Šuligoj T, Vaux L et al. Unravelling the specificity and mechanism of sialic acid recognition by the gut symbiont Ruminococcus gnavus. Nat Commun 2017;8:2196.
Tailford LE, Owen CD, Walshaw J, Crost EH, Hardy-Goddard J, Le Gall G, de Vos WM, Taylor GL, Juge N. Discovery of intramolecular trans-sialidases in human gut microbiota suggests novel mechanisms of mucosal adaptation. Nat Commun 2015;6:7624.
Juge N, Tailford L, Owen CD. Sialidases from gut bacteria: a mini-review. Biochem Soc Trans 2016;44:166–175.

Andrew Bell
Quadram Institute Bioscience, Norwich Research Park, NR4 7UQ, UK
Andrew Bell is a post graduate student in the Department Gut Microbes and Health at the Quadram Institute studying the impact of a novel mechanism of sialic acid metabolism on gut homeostasis. He obtained his BSc in Biochemistry from the University of East Anglia and spent 2 years working at the John Innes Centre on chlorophyll metabolism in pea (Pisum sativum) before joining Nathalie Juge’s group at the Quadram Institute in 2015 for his doctoral work.

Emmanuele Severi
Department of Biology, University of York, Wentworth Way, York YO10 5DD, UK
Emmanuele Severi is a post-doctoral research associate in the Department of Biology at the University of York working in the laboratory of Prof Gavin Thomas. He has worked on sialic acid transport and metabolism at different times in his career, and has been a member of the Society since 2002.

Nathalie Juge
Quadram Institute Bioscience, Norwich Research Park, NR4 7UQ, UK
Nathalie Juge is a Research Leader at the Quadram Institute Bioscience (QIB) in Norwich, Deputy Leader of the Institute Strategic Programme, the Gut Microbes and Health (GMH), and Honorary Professor at the School of Biological Sciences, University of East Anglia. She leads a Research Group on the glycobiology of host-microbe interactions in the gut with a focus on mucin-derived sialic acid metabolism by the gut microbiota (https://quadram.ac.uk/nathalie-juge).

Gavin H. Thomas
Department of Biology, University of York, Wentworth Way, York YO10 5DD, UK
[email protected]
@GavinHThomas
Gavin H. Thomas is a Prof. of Microbiology in the Department of Biology at the University of York and have worked on bacterial sialic acid transport and catabolism for over a decade. He has been a member of the Society since 1995, was Editor of Microbiology Today (2003–2006), serving on Council in the same period, served on Communications committee (2012–2015) and the Equality & Diversity committee (2013–2015). He has been an Editor, Senior Editor and is now Deputy Editor-in-Chief of the Society’s leading journal Microbiology (http://thomaslabyork.weebly.com).
Why does microbiology matter?
Gavin: It matters for our health – only now are we really starting to understand the molecular composition and function of our microbiome, which likely impacts on our body in multitudinous ways.
What advice would you give to someone starting out in this field?
Gavin: Read widely and beyond ‘your bug’ – microbes can do pretty much everything and have usually invented multiple different ways to do it!
Images: H3N2 influenza virus particles. CDC / Science Photo Library.
Fig. 1. Andrew Bell.
Fig. 2. Emmanuele Severi.
