Foodborne diseases: sequencing for answers
Issue: Microbes and Food
07 August 2018 article
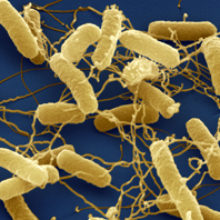
‘Food poisoning’, ‘stomach bug’ and ‘gastroenteritis’ are some of the ways in which we commonly refer to foodborne diseases, which are a constant challenge for humans. We are all susceptible to the bacteria that cause foodborne diseases, but the more vulnerable in society – the young, old, immunocompromised and those with limited resources and infrastructure – often suffer the gravest consequences.
While most of us experience mild and self-resolving diarrhoea with some nausea and vomiting for a few days, foodborne diseases can lead to severe, life-threatening conditions such as kidney failure and neurological disorders. Foodborne diseases affect lives and economies.
Be smart, be safe
Among the most common causes of foodborne diseases are Salmonella, Escherichia coli and Campylobacter. These bacteria are ubiquitous in the environment and typically reside within the intestines of farmed animals like cows, pigs and chickens and within unmanaged wildlife. During slaughter and processing, bacteria can contaminate equipment and carcasses. A good example of this is the topical contamination of chicken by Campylobacter. Other bacteria are found deep in tissues; for example, Salmonella can survive within the peripheral lymph nodes of cows and pigs. These lymph nodes are small and challenging to remove during processing and often get incorporated into minced meat. Improper handling and consumption of contaminated food leads to food poisoning but simple measures, such as not washing raw chicken to prevent Campylobacter contamination of our kitchens and thorough cooking of mince to eliminate Salmonella, can be highly effective in reducing our risk. The Food Standards Agency ensures that products are appropriately labelled with storage and handling advice to make food safer for consumers. However, we can also be exposed to zoonotic bacteria via direct contact with infected people and animals, so proper handwashing is an essential part of preventing infection.
Animals can suffer too
There is a common perception that foodborne pathogens are harmless to their animal reservoirs, but this is not necessarily true. Some types of Salmonella cause transient diarrhoea in cows and pigs, while others can cause typhoid-like disease and lead to abortion. Even Campylobacter, which was long-regarded as a harmless commensal of chickens, has been associated with disease and pathology in some broiler lines. Vaccination is an attractive option to reduce the burden of foodborne diseases. However, there are currently no licensed vaccines for Campylobacter in chickens or Shiga toxin-producing E. coli in cattle. Some Salmonella vaccines are available for farmed animals, but how they confer protection is not well understood. In concert with vaccines, better hygiene and changes in farm practices can also help to limit foodborne pathogens, as evidenced by the National Control Plan for Salmonella in poultry in the UK. Given the rise of drug-resistant superbugs, we cannot simply eradicate foodborne pathogens with antimicrobials. How then can we make food safer and improve animal welfare?
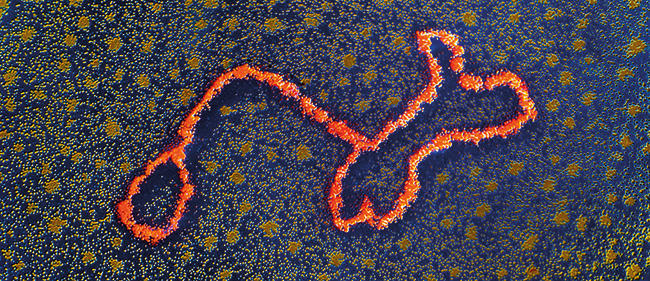
From sequence to infection
Researchers have had access to the genetic blueprints of key foodborne pathogens for decades, but bacterial genomes are complex. It can be difficult to predict which genes contribute to disease, the impact of sequence variation and which genes would make good targets for vaccines and other interventions. Due to advances in sequencing technologies, the number of bacterial genome sequences in databases is increasing exponentially, but relatively few studies have sought to link bacterial genotype and phenotype during infection.
Foodborne bacteria are relatively easy to study. We can grow them in large numbers in the laboratory and have numerous molecular methods to genetically manipulate them. For example, we can knock-out genes to study their function, or knock-in nucleotide markers to make bacteria trackable, and we are constantly developing more sophisticated ways to study them. However, it is still unrealistic to study bacterial pathogenesis gene-by-gene or strain-by-strain, especially in relevant animal species, owing to the large number of animals, time and costs involved. To overcome these obstacles, researchers have developed clever approaches to study foodborne diseases with minimal animal use.
Sequencing approaches have been invaluable to the study of all aspects of foodborne diseases, from outbreak investigations and epidemiology, to pathogenesis and evolution. The prediction of virulence and zoonotic risk of foodborne pathogens remains a key challenge of the post-genomic era, but novel methods are aiming to tackle these. A method called transposon directed insertion sequencing (TraDIS) combines bacterial mutagenesis with deep sequencing, to identify genes which are essential for pathogen survival. By encouraging a mobile genetic element called a transposon to insert into genes, gene function is disrupted. Then, using sequencing, disrupted genes can be identified, and the number of bacteria with a defined disruption present in a population can be quantified. If a gene disruption leads to reduced bacterial survival, we can infer that that gene is important for survival. In this way, we have found out which genes Salmonella needs for bile tolerance, a requirement for intestinal bacteria. We also studied the roles of over half the genes of Salmonella during colonisation of intestines of chickens, pigs and calves. Interestingly, some genes were essential to colonise all the reservoir hosts, which would make them good targets for vaccines, but others were host-specific, suggesting that Salmonella uses different strategies to survive in them. Similar studies have also now defined conditional essential genes of E. coli and Campylobacter.
Along with understanding gene function during infection, it is also important to know which bacterial strains pose greater risk, but this is difficult. There are over 2,600 serovars (distinct variations within a species distinguished by their cell surface antigens) of Salmonella: some cause disease in humans, some in animals and some in both. So how do we decide which are high-risk? To try to answer this, a new method to simultaneously test multiple serovars was developed. All Salmonella serovars share genes which vary in their sequence between serovars. These variations are stable, serovar-specific and can be identified by sequencing, to determine which serovars are present in a mixed population and at what abundance. From the presence or absence and abundance of serovars under different conditions, we can infer serovar fitness. Using this method, we studied 11 serovars in 10 tissues of cows simultaneously and found that all of them could survive within peripheral lymph nodes, despite varying in their capacity to colonise the intestines, providing valuable information for vaccine design. Sequence analysis can also be used to predict whether some bacterial strains are more zoonotic than others. Using machine-learning algorithms to analyse genome sequences of hundreds and thousands of bacterial isolates from humans and animals, computers can be trained to differentiate between them and group them based on their genetic traits. When this was applied to E. coli and Salmonella, a subset of bovine strains with human-like traits and vice versa were identified, suggesting that they pose a risk of zoonosis. Different sequencing-based approaches are also being used to link the presence of bacterial genes with traits of interest and show promise in identifying bacterial genes that may influence zoonosis.
Closing the gap
Bacterial foodborne pathogens remain a major threat to humans and animals alike, but using different sequencing and computational methods, we are slowly beginning to unravel the mysteries of these masters of misery. We now need to use the knowledge we have acquired, to develop effective drugs and vaccines to control these pathogens and reduce the burden of foodborne diseases. Long-term research is required to bring such strategies to fruition. In the meantime, we must remain cautious. Surveillance and prevention are essential while we await solutions.
Further reading
Chaudhuri R.R. & others (2013). Comprehensive assignment of roles for Salmonella Typhimurium genes in intestinal colonization of food-producing animals. PLoS Genet 9, e1003456. doi:10.1371/journal.pgen.1003456
Langridge G.C. & others (2009). Simultaneous assay of every Salmonella Typhi gene using one million transposon mutants. Genome Res 19, 2308–2316. doi:10.1101/gr.097097.109
Lupolova N. & others (2016). Support vector machine applied to predict the zoonotic potential of E. coli O157 cattle isolates. Proc Nat Acad Sci 113, 11312–11317. doi:10.1073/pnas.1606567113
Vohra P. & others (2018). Quantifying the survival of multiple Salmonella enterica serovars in vivo via massively-parallel whole genome sequencing to predict zoonotic risk. Appl Environ Microbiol 84, e02262–17. doi:10.1128/AEM.02262-17
Yahara K. & others (2017). Genome-wide association of functional traits linked with Campylobacter jejuni survival from farm to fork. Environ Microbiol 19, 361–380. doi: 10.1111/1462-2920.13628

Prerna Vohra
The Roslin Institute & Royal (Dick) School of Veterinary Studies, University of Edinburgh, Easter Bush, Midlothian EH25 9RG
[email protected]
Twitter: @DrPrernaVohra
Prerna Vohra is a postdoctoral researcher with a keen interest in enteric pathogens and how they interact with their hosts. She began her research career by studying Clostridium difficile during her MSc and PhD. She then moved on to vaccine-related projects, first looking at Bacteroides fragilis as a delivery system for vaccines for diarrhoeal diseases, and later trying to develop novel glycoconjugate vaccines for veterinary pathogens. Her current research focuses on understanding the pathogenesis of Salmonella and Campylobacter in cattle and chickens, respectively, to inform the design of vaccines or control strategies. She has been a member of the Microbiology Society since 2007 and is an ECM Forum member.
What do you love most about your job?
I love learning new things and my job gives me ample opportunities for this. I love that I can be creative to find new ways to solve problems. I also love the diversity of my day-to-day work. It can be challenging, but that’s what makes it fun.
What skills are required in your position on a day to day basis?
I use my lab skills – bacteriology, molecular microbiology, immunology and in vivo skills – to do my research; computational skills – data organisation, bioinformatics and statistical analysis – to understand the results; and writing and presentation skills to communicate the outcomes, along with project planning and time management of course!
Images: Salmonella typhimurium bacteria, coloured scanning electron micrograph (SEM). Juergen Berger/Science Photo Library.
False-colour transmission electron micrograph of a DNA transposable genetic element or ‘transposon’, seen here forming a characteristic stem-and-loop structure. Professor Stanley Cohen/Science Photo Library.
