Influenza pseudotype research and development
Issue: Microbial Tools
15 May 2018 article
Undertaking research and development (R&D) on novel therapeutics and vaccines for emerging viral pathogens is fraught with numerous obstacles, one of which is the requirement for high- containment laboratory facilities.
We and others have pioneered the design, construction and deployment of retroviral pseudotype viruses (PV) that are now having significant impact on preclinical research undertaken by pharma, and by public and animal health institutes worldwide. These pseudotypes are biosafe, replication-defective retroviruses and lentiviruses that have been repurposed from gene therapy applications. They have heterologous (foreign) viral glycoproteins on their surface that originate from the viral pathogen. They thus mimic the wild- type virus with respect to cell entry, but can be handled by end users in biosafety level (BSL)-1 instead of BSL-3/4 laboratories. They have been described elegantly by Robin Weiss FRS (a former President of the Microbiology Society) as ‘sheep in wolves’ clothing’. These incorporate a quantifiable reporter, usually firefly luciferase or GFP (which is integrated into the target cell genome), enabling sensitive and high-throughput cell entry-based assays to be performed. The pseudotype-based assays are simple to upgrade, as only the nucleotide sequence of the surface glycoprotein(s) is required. These sequences are readily obtained from NCBI and GISAID. This makes them particularly ideal for emerging RNA viruses, e.g. avian influenza, rabies, SARS/MERS coronaviruses, Ebola. These pseudotype viruses are very stable over a range of temperatures and can be freeze-dried for efficient deployment to end users. It is hoped that this will facilitate the roll-out of such assays to low- and middle-income countries (LMIC). The virus neutralisation assays performed using pseudotype viruses are both serum and antigen sparing, and enable the study of functional responses, which correlate strongly with those measured using cognate wild-type virus assays. This results in significant cost saving for the end user, given the exorbitant infrastructure and staff costs of undertaking serological assays with high-containment viruses of human and animal importance.
Viral Pseudotype Unit
Expertise at the University of Kent in pseudotype technology led to the creation, in 2011, of the Viral Pseudotype Unit (VPU). The VPU acts as an interface between academia, industry, and animal and public health laboratories, with the purpose of translating basic virus research into in vitro cell culture pseudotype-based assays that can be readily employed for the characterisation of vaccines, antivirals and therapeutic antibodies. This has led to the use of pseudotype assays in the following broad R&D areas: immunogenicity testing of current and preclinical viral vaccines including new ‘universal’ vaccine prototypes; screening of monoclonal antibodies to isolate those that broadly neutralise across multiple subtypes/species of the same virus; characterisation of antibody standards for serological assays; study of virus restriction factors (entry); and drug screening/resistance studies.
Functional serological assays to measure antibody responses against the haemagglutinin (HA) globular head, HA stalk and neuraminidase (NA)
Influenza pseudotypes have been used to undertake serological assays for big pharma and biotech clients, and will hopefully contribute one day towards the licensure of influenza vaccines, at least as an adjunct assay. In a joint study with Novartis Vaccines, we showed that pseudotypes could be used to quantify neutralising antibody responses elicited by a pre-pandemic H5N1 vaccine and that these correlated significantly with those measured by haemagglutination inhibition (HI), single radial haemolysis (SRH) and microneutralisation. Pseudotype neutralisation assays for influenza have been shown to be exquisitely sensitive for the measurement of responses to the HA stalk, which is one of the primary targets of many big pharma ‘universal vaccine’ approaches. The responses directed against the stalk can be separated from those against the head by making use of a hybrid HA pseudotype. One such virus that we have created has an HA stalk derived from the 2009 pandemic H1N1 strain, and an HA globular head derived from an H11 strain which is found in wild birds. Humans have negligible serological reactivity to H11, so if a neutralisation assay is undertaken with this hybrid HA pseudotype, one can readily measure the response against the HA stalk only. The traditional HI assay (for which a correlate of immunity exists) used by the regulators only measures responses against the globular HA head and thus is not fit for purpose for the licensing of many new ‘universal vaccines’. HA stalk serological assays, such as those based on pseudotypes or ELISA, will no doubt gain prominence as many of these new vaccines move down the clinical pipeline.
Recently, a pseudotype-based enzyme-linked lectin assay (PV- ELLA) has been developed, which allows the quantification of antibody responses against NA. This assay innovatively uses NA-only pseudotypes as a source of NA. The availability of this PV-ELLA to the R&D community is instrumental as it means that ELLA assays can be performed without requiring access to reverse genetic (RG) viruses, which often have IP attached. Also, there are moves towards standardising the amount of NA in recombinant vaccines, making PV-ELLA assays increasingly important.
Pseudotypes are also gaining acceptance as a method for the identification of antibody standards that can be used to harmonise serological outputs across multiple different assays. This was recently undertaken for the Ebola virus by the NIBSC.
Characterisation of therapeutic mAbs
The influenza HA pseudotype neutralisation assays and know-how have been used in pivotal collaborations with pharma companies that isolate monoclonal antibodies derived from individuals whose immune systems have successfully responded to influenza infection. One of these monoclonals, FI6 developed by Humabs in Switzerland, targets the fusion peptide on the HA stalk and is thus able to protect against all subtypes of influenza and therefore critical for informing universal vaccine design. Pseudotype viruses were instrumental in the characterisation of this broadly neutralising antibody. The availability of pseudotype viruses to industry has resulted in significant time/cost savings and increases in assay sensitivity and ultimately productivity, and has been moved forward into other pipelines for rabies, Ebola and MERS.
New viruses, new reservoirs
Not all influenza A subtypes have their reservoir in wild birds. Two new influenza subtypes, H17 and H18, have been discovered in bats, and the potential for spillover into humans or intermediate hosts is unknown at the present time. However, using our pseudotype technology, we have been able to construct an H17N10 pseudotype and shown that this virus (unlike H1–H16) does not use sialic acid as its receptor. Research is ongoing in our and other laboratories to identify this receptor. We additionally showed that H17N10 can be efficiently neutralised by the mAb FI6, and also by serum from individuals who have a high antibody response against the H1 stalk (as measured using the hybrid HA pseudotype assay).
In conclusion, I would say that the future is bright for using pseudotypes in virology R&D. As surrogates of the wild type, they offer significant safety advantages, and their readily adaptable and upgradable nature allows them to be used to study multiple targets of the antibody response.
Further reading
Biuso, F. & others (2017). The study of antibody responses to influenza neuraminidase using a lentiviral pseudotype based ELLA. bioRxiv (preprint). doi:10.1101/218800
Carnell, G. W. & others (2015). Pseudotype-based neutralization assays for influenza: a systematic analysis. Front Immunol 6, 161. doi:10.3389/fimmu.2015.00161
Ferrara F. & others. (2015). The application of pseudotypes to influenza pandemic preparedness. Future Virol 10, 731–749. doi:10.2217/fvl.15.36
Temperton, N. J. & others (2015). Retroviral Pseudotypes – From Scientific Tools to Clinical Utility. In eLS, John Wiley & Sons, Ltd (Ed.). doi:10.1002/9780470015902.a0021549.pub2
Temperton, N. (2016). The Viral Pseudotype Unit: viral pseudotype R&D, dissemination and education. Future Virol 11, 113–116. doi:10.2217/fvl.15.109

Nigel Temperton
Medway School of Pharmacy, Universities of Greenwich and Kent at Medway
[email protected]
https://orcid.org/0000-0002-7978-3815
Nigel Temperton’s research interests lie primarily in emerging/re-emerging and transboundary viruses (SARS/MERS coronavirus, pandemic/inter-pandemic influenza, and Ebola), and the methods for their control – in particular, control by vaccination-induced antibody responses. He has established novel virus antibody neutralisation assays for these high-containment viruses by exploiting retroviral and lentiviral vector technologies.
Nigel has been a member of the Microbiology Society since 2003. The Society has contributed significantly towards the impact and reach of his group’s pseudotype work by giving platforms for PhD students to present their work to a wide spectrum of microbiologists.
How did you enter this field?
I entered this field during my time as a Wellcome Trust-funded postdoc studying endogenous retroviruses in Robin Weiss’ group at UCL. After a lunchtime seminar given by a Hong Kong academic visitor who had himself been infected by the SARS coronavirus, Robin said to me “Nigel, what you need to do is develop a SARS pseudotype so we can do some serology”. The rest is history. I have dedicated my career to exploiting these pseudotypes ever since.
What is the most rewarding part of your job?
The most rewarding part of my job is first and foremost the nurturing and training of the next generation of ‘pseudotypers’. I also enjoy promoting pseudotype technologies by any means available, including over social media @ViralPseudotype.
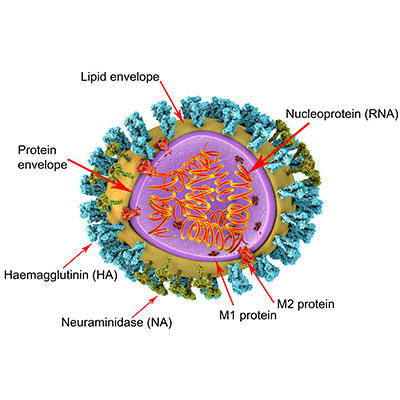
Images: Coloured transmission electron micrograph of swine influenza (H1N1) virus particles (virions) determined to be the cause of the April 2009 outbreak (originated in Mexico City, Mexico). Dennis Kunkel Microscopy/Science Photo Library. 3D illustration of influenza virus particle structure. The virus consists of a ribonucleic acid (RNA, orange coils) core, surrounded by a nucleocapsid (purple) and a lipid envelope (yellow). Spanning the capsid and envelope are M2 proteins (red) that act as proton pumps. In the envelope are two types of protein spike, haemagglutinin (H, blue) and neuraminidase (N, green), which determine the strain of virus. Kateryna Kon/Science Photo Library.
