Microbe Machines: How ‘souped-up’ bacterial motors produce more torque
Posted on May 17, 2016 by Anand Jagatia
Many bacteria have tiny motors inside them, which they use to zip around like miniature submarines. Recently, scientists have managed to image a diverse selection of these nanomachines in more detail than ever before, and gain fundamental insights into how they work.
Bacterial motors are an incredible feat of evolutionary engineering. The motor in E. coli, for example, spins 125 times a second at room temperature, and pound for pound it produces the same amount of power as a turboprop aeroplane engine. Even more amazingly, these complex machines seamlessly self-assemble in a precise order, piece by piece, from around 20 different proteins and 50 different genes.
Bacteria use these motors to rotate corkscrew-shaped filaments called flagella, which act like propellors and move the cell forward. The motor’s powerhouse is embedded in the bacterium’s cell membrane, and it drives the rotation of the flagellum by converting the movement of charged particles (usually protons) into a turning force, or torque.
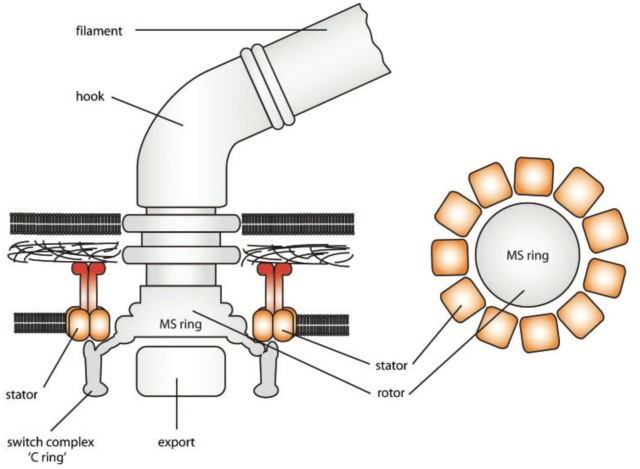
The diagram below shows how a bacterium makes this happen. Motors are made up of a rotating component, called the rotor, and a stationary component, called the stator. In the flagellar motor, protons flow through channels in a ring of ‘stator complexes’ that form the stator. The stators use the energy from the flowing protons to exert force on the rotor, which in turn transmits torque to the filament – propelling the bacterium through its environment.
Petrified bacteria
Observing individual motors inside bacteria is extremely difficult, because of how small and complex they are. But now, a team of researchers has been able to view the functional heart of motors from different bacteria in unprecedented detail, using a rather ingenuous technique.
First, an entire bacterium is plunged into a liquid mixture that has been cooled to around -200ºC by liquid nitrogen. Normally, water in the bacterium would expand as it freezes, rupturing the cell, but here the process occurs so quickly that ice crystals don’t have a chance to form.
“We place this petrified bacterium into an electron microscope to get an image,” explains Morgan Beeby from Imperial College London, who led the study. “We tilt the image over a range of angles, and then just like a CT scan we use this set of 2D images to create a 3D model.”
You can then take a slice through this 3D model to get a cross sectional image of the bacterial motor. Because the motor is so small, however, the quality of the resultant image is rather fuzzy and contains a lot of noise. The researchers get round this problem using a clever trick – by imaging hundreds of genetically identical bacteria and making a composite average, it’s possible to ‘wash out’ the noise and clarify the details, to form a much cleaner image (compare C and F in the image below).
Image credit: Beeby et al 2016, PNAS
This is a revolutionary technique (no pun intended) as it allows scientists to see the molecular machinery as it actually exists in the cell. In fact, the motors used to make the image above were probably rotating at the time they were frozen.
The team imaged a range of motors inside different species and were surprised to find that, even though the motors were all based on the same basic motor ‘chassis’, there were striking differences.
“We could see that the motors all had a common core that looks like the Salmonella motor, but a lot of them had additional bits bolted on,” says Morgan. “We wanted to understand why this was, so we looked at motors in three bacteria that have very different swimming abilities: Campylobacter, Vibrio and Salmonella.”
Swimming through treacle
Biophysicists have shown that flagellar motors in different bacteria are able to produce different amounts of torque, and this affects how well they are to swim in thick, sticky liquids. For example, the gut bacterium Campylobacter (which can cause food poisoning), can bore through the viscous mucous lining the intestines, whereas other bacteria like Salmonella and Vibrio would be stopped in their tracks.
Salmonella bacteria are stopped in their tracks by viscous fluid, but Campylobacter are able to churn their way through.
Video courtesy of the researchers
So how are some bacteria able to generate this extra torque? The mechanism has so far remained unknown, in part because the motors themselves are so hard to visualise. But Morgan’s team noticed that in bacteria with higher torque motors – like Campylobacter – the stator complexes that power the machine were both more numerous and positioned in a wider circle around the central rotor ring. It’s this combination that allows some bacterial motors to produce so much force.
Spanners in the works
“If you have a long spanner you can generate more torque than you can by using a short spanner,” explains Morgan. “You have to move the spanner through a greater distance, so it’s a trade-off, but you produce more torque. So we thought, maybe by positioning more stators further away from the rotors in a wider ring, the bacteria are exerting greater force.”
When they crunched the numbers, the team showed that their calculations for torque, based on the number of stator complexes and the distance from the rotor, matched up consistently with real world values for motor torque in different species.
“We’ve been able to make an important contribution to understanding the biological mechanisms of the motor,” says Morgan. “The great thing is that the images of these structures allowed us to rationalise the different levels of torque that people have been seeing for years, but no one knew why.”
The researchers think that the other proteins in the images, which form complex structures that differ between species, have evolved to scaffold the stator complexes further out from the rotors. This has enabled bacterial species to adapt their motors to different environments – from freshwater to gut mucus – so that they can seek out more favourable conditions or escape danger.
After all, when you’re a bacterium, it’s sink or swim.

