Using insect larvae to study disease
Posted on May 22, 2019 by Gerard Sheehan
Researchers from Maynooth University have been using larvae as a model to examine the progress of Staphylococcus aureus infection. Here, PhD student Gerard Sheehan discusses his research which has recently been published in the journal Microbiology.

I am currently in my third and final year as a PhD student in the Medical Mycology lab at Maynooth University, Ireland. I would describe my PhD as quite broad; encompassing areas of microbiology such as antibiotic resistance, bacterial and fungal pathogenesis and modelling of infection processes in an alternative animal model (larvae of the insect Galleria mellonella).
G. mellonella larvae have been used over the past 20 years to study the virulence of microbial species because they possess many advantages over other in vivo systems. They are low-cost, easy to use, free from legal/ethical restrictions and generate results within hours or days. Most importantly, they produce results similar to those obtained using mice. This is because the insect immune system is up to 90% similar to our own innate immune response. We believe larvae can be used to study infection processes and pathogenesis and my PhD provides evidence for this.
In our study, we focus on Staphylococcus aureus and look at how it infects larvae, examine the extent of the cellular immune response, characterise dissemination of the bacterium throughout the host and also use proteomics to detail the humoral immune response to S. aureus.
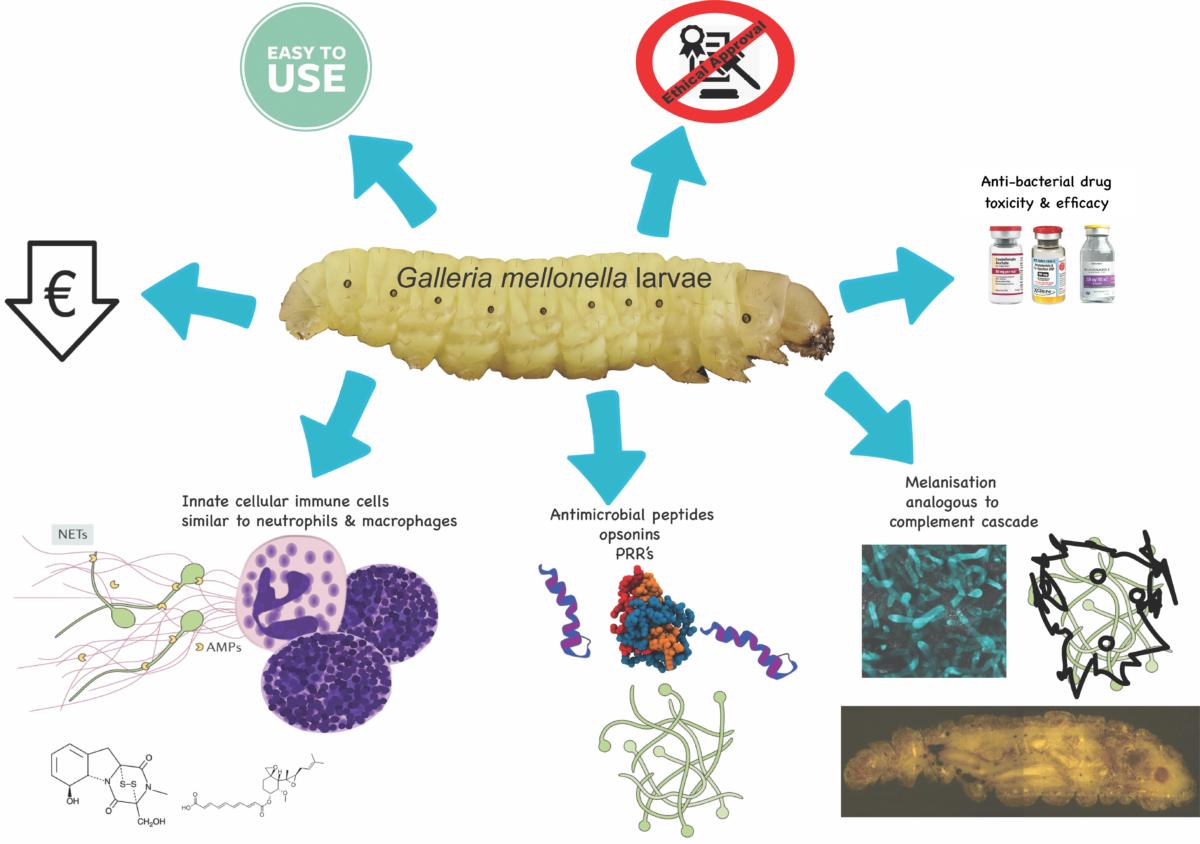
Advantages associated with using G. mellonella larvae.
In our lab we use a lot of proteomics which means plenty of bioinformatical analysis. Advancements in technology mean we can detect thousands of proteins in a complex sample, and this gives us an idea of what exactly is going on during the infection.
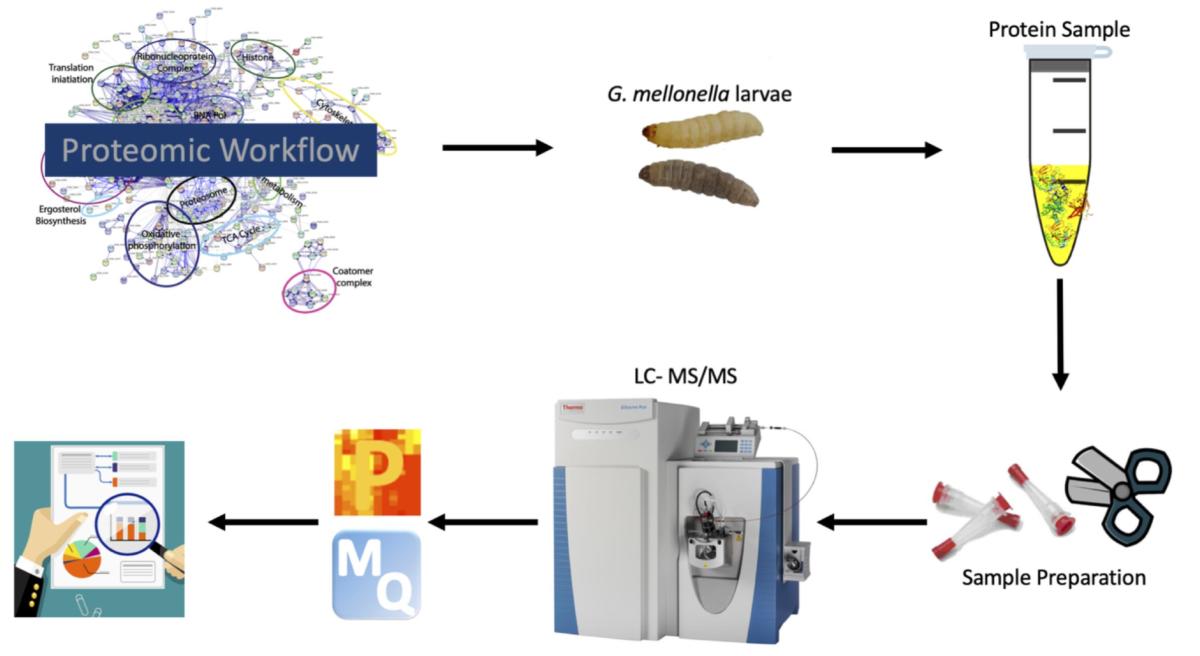
Workflow for proteomics. Hemolymph (equivalent to human blood) is extracted from S. aureus-infected larvae. Protein is then purified and trypsin digested to generate peptides, and these are subjected to liquid chromatography–mass spectrometry. The data is analysed by a range of statistical packages (e.g. MaxQuant, Perseus).
Our results demonstrate that S. aureus kills larvae in a dose-dependent manner, interacts with and overcomes the cellular immune response and produces structures similar to abscesses which form during human infection. Interestingly, we use Cryoviz-imaging to look inside the larvae during different timepoints following infection. The Cryoviz imaging system was originally developed to image internal structures of mice, but we have adapted it to use with larvae. Larvae are placed in an embedding compound and frozen. Samples are mounted on the stage and the machine cuts 10 μm slices. It images after each slice which allows us to visualise the internal organs of the larvae. The entire process can be carried out in one day. From this, we can visualise abscess formation but also extensive melanisation (similar to inflammation) which is a protective response in insects.
From the proteomic results we found proteins such as peptidoglycan-recognition protein LB and peptidoglycan-recognition-like protein B were increased five- fold at six hours post infection and these play a role in early recognition of S. aureus. By 24 hours after infection, there was an increase in the abundance of a range of antimicrobial peptides/proteins, such as gloverin (86-fold), cecropin-D (30-fold) and lysozyme (18-fold) which display in vivo anti-bacterial activity. We also identify a range of candidate proteins which may have a role in abscess formation in larvae.
We then discuss the results in terms of S. aureus infection in mammals and find many similarities (such as rapid bacterial proliferation, nodule formation, alteration in immune cell number, increased antimicrobial peptide abundance).
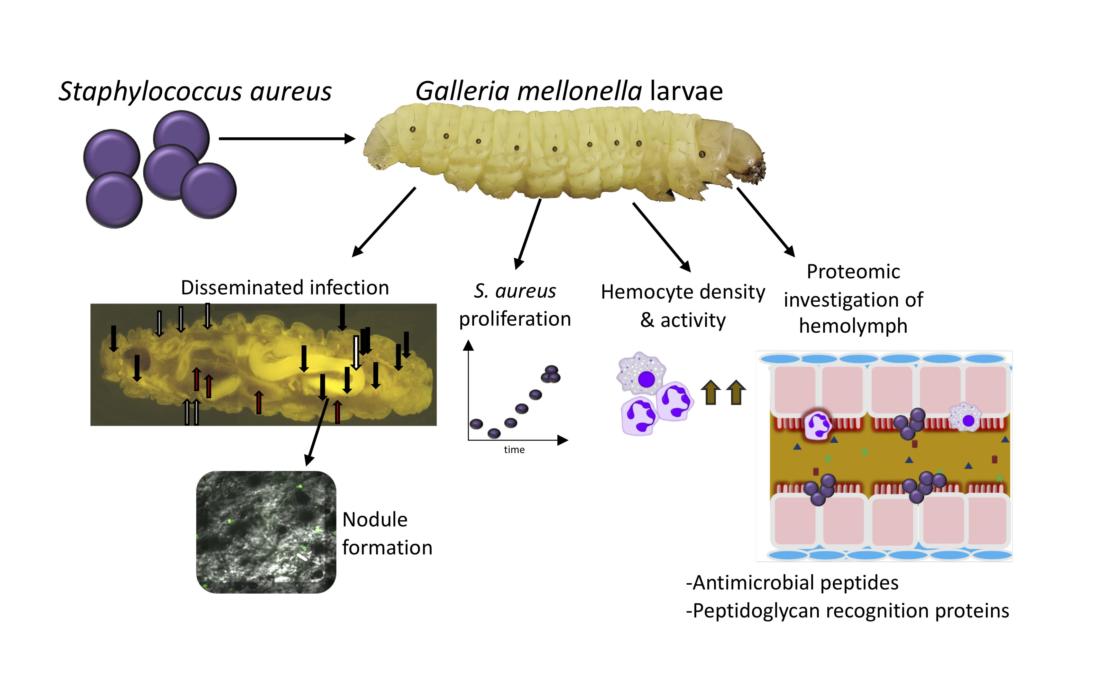
Summary of the results obtained in this study. Infection by S. aureus results in dissemination of the bacterium throughout the host and changes in the cellular and human immune response.
These results demonstrate that larvae are a good model to study the pathogenesis of S. aureus within its host and the immune responses activated to defend against its development. We hope this may be expanded to other bacteria and allow the use of larvae to study the efficacy and interaction of novel antibiotics with S. aureus and the innate immune response.