Schoolzone: The nitrogen cycle
08 May 2012

Various processes are responsible for recycling the chemicals necessary for life on Earth. The nitrogen cycle is the most complex of these. Carbon, sulfur and phosphorus are the other main cycles. In this article we explore how nitrogen is cycled and the important role of microbes in this cycle.
Nitrogen is required by all living organisms for the synthesis of organic molecules such as amino acids, nucleic acids and proteins. The Earth’s atmosphere contains almost 80% nitrogen gas. It cannot be used in this form by most living organisms until it has been fixed, that is reduced (combined with hydrogen), to ammonia. Green plants, the main producers of organic matter, use this supply of fixed nitrogen to make proteins that enter and pass through the food chain. Micro-organisms (the decomposers) break down the proteins in excretions and dead organisms, releasing ammonium ions. These two processes form part of the nitrogen cycle.
The nitrogen cycle
The nitrogen cycle is the movement of nitrogen between the earth and the atmosphere. It consists of a series of processes that convert nitrogen gas to organic substances and these back to nitrogen in nature. It is a continuous cycle maintained by the decomposers and other bacteria. The nitrogen cycle can be broken down into four types of reaction and micro-organisms play roles in all of these (see Table 1).
Nitrogen fixation
Nitrogen gas is composed of two atoms of nitrogen linked by a very strong triple bond. This makes it chemically unreactive and large amounts of energy are required to break the bond. Nitrogen gas can be fixed in three ways.
- Atmospheric fixation. This occurs spontaneously by lightning; only a small amount (5–8%) is fixed this way. Lightning allows nitrogen and oxygen to combine to produce various oxides of nitrogen. These are carried by the rain into the soil where they can be used by plants.
- Industrial fixation. The Haber–Bosch process is used to make nitrogen-containing fertilizers. This is a very energy-inefficient process.
- Biological fixation. Nitrogen-fixing bacteria fix 60% of nitrogen gas in the atmosphere.
Biological fixation
The reduction of nitrogen gas to ammonia is energy-intensive. It requires 16 molecules of ATP and a complex set of enzymes to break the bonds so that the nitrogen can combine with hydrogen. Its reduction can be written as:
N2 + 3H2→ 2NH3
Relatively few bacteria (the nitrogen-fixing bacteria) are able to carry out this reaction. Fixed nitrogen is made available to plants by the death and lysis of free-living nitrogen-fixing bacteria or from the symbiotic association of some nitrogen-fixing bacteria with plants.
Table 1. Reactions of the nitrogen cycle.
| Reaction | Micro-organism | Conditions | Process |
| Nitrogen fixation | Nitrogen-fixing bacteria, e.g. Rhizobium | Aerobic/anaerobic | The first step in the synthesis of virtually all nitrogenous compounds. Nitrogen gas is fixed into forms other organisms can use. |
| Ammonification (decay) | Ammonifying bacteria (decomposers) | Aerobic/anaerobic | The decomposers, certain soil bacteria and fungi, break down proteins in dead organisms and animal wastes, releasing ammonium ions which can be converted to other nitrogen compounds. |
| Nitrification | Nitrifying bacteria, e.g. Nitrosomonas, Nitrobacter | Aerobic | Nitrification is a two-step process. Ammonia or ammonium ions are oxidized first to nitrites and then to nitrates, which is the form most usable by plant. |
| Denitrification | Denitrifying bacteria | Anaerobic | Nitrates are reduced to nitrogen gas, returning nitrogen to the air and completing the cycle. |
Types of nitrogen-fixing bacteria
Some nitrogen-fixing bacteria are free-living in the soil, fixing nitrogen independently of other organisms, e.g. Azotobacter (aerobic) and Clostridium (anaerobic). Other nitrogen-fixing bacteria form symbiotic associations with plants.
Root-nodulated legumes, such as peas and beans, with e.g. Rhizobium. Free-living rhizobia invade the legume through an infection thread formed in the root hair of the plant. The infection thread is constructed by the root cells, not the bacteria, and is formed only in response to infection. The infection thread grows through the root hair cells and penetrates other root cells nearby, often with branching of the thread. The root cells then proliferate to form a root nodule.
Within a week of infection, small nodules are visible to the naked eye. Each root nodule is packed with thousands of living Rhizobium bacteria (known as bacteroids).
Root-nodulated non-legumes, a diverse group of woody species such as alder, with e.g. Frankia. These filamentous bacteria infect the roots of plants forming actinorhizal root nodules.
Azolla (tiny free-floating water ferns) with e.g. Anabaena azollae. This is a cyanobacterium that infects new leaves of Azollas as they develop from the stem. Strings of Anabaena get caught in tiny leaf hairs that grow from a dimple on the developing leaf. The dimple grows larger into a pouch-like structure that eventually closes up, locking the Anabaena inside the leaf.
Adapting to their environment
Nitrogen-fixing bacteria contain an enzyme complex called nitrogenase which catalyses the conversion of nitrogen gas to ammonia. It supplies hydrogen ions as well as energy from ATP. The nitrogenase complex is sensitive to oxygen, becoming inactivated when exposed to it. This is not a problem with the free-living anaerobic bacteria such as Clostridium. Free-living aerobic bacteria have a variety of different mechanisms for protecting the nitrogenase complex, including high rates of metabolism and physical barriers. Azotobacter overcome this problem by having the highest rate of respiration of any organism, thus maintaining a low level of oxygen in their cells.
Rhizobium contains leghaemoglobin. This functions similarly to haemoglobin, i.e. it binds to oxygen. This provides sufficient oxygen for the metabolic functions of the bacteroids, but prevents the accumulation of free oxygen that would destroy the activity of nitrogenase.
Frankia and Anabaena are able to exclude oxygen by carrying out the fixation in specialized structures known, respectively, as a vesicle and a heterocyst. The thick walls of the vesicle and heterocyst form an oxygen diffusion barrier.
Nitrification
This is the oxidation of ammonium compounds to nitrites and then to nitrates by the nitrifying bacteria. During these oxidation reactions energy is released. The nitrifying bacteria are chemoautotrophs and are able to use this source of energy to produce organic compounds from inorganic ones (photo-autotrophs use light energy to produce organic compounds from inorganic ones).
Nitrification is a two-step process.
- Bacteria of the genus Nitrosomonas convert ammonium ions to nitrites (NO2–). (Nitrite is toxic to plants and animals in high concentrations.)
- Bacteria of the genus Nitrobacter convert nitrites to nitrates (NO3–). The nitrates can then be taken in by plants.
Nitrification occurs in well-drained and aerated soils at neutral pH.
Denitrification
This is the conversion of nitrates into primarily nitrogen gas, but also nitrous oxide gas by the denitrifying bacteria, e.g. Pseudomonas. Denitrifying bacteria transform nitrate in extremely wet soils and swampy grounds where there is very little oxygen, i.e. the conditions are anaerobic. The bacteria get the oxygen they need for respiration from the breakdown of nitrates. The gases that are formed escape into the atmosphere completing the nitrogen cycle. This can be a harmful process as fixed nitrogen is removed from the soil making it less fertile.
Ammonification (decay)
This is the conversion of organic forms of nitrogen (e.g. in dead organisms and their excretions) into inorganic nitrogen. A wide range of soil fungi and bacteria, called the decomposers, carry out the ammonification process. The decomposers consume the organic matter, and the nitrogen contained in the dead organism is converted to ammonium ions. The ammonium is then converted to nitrates by the nitrifying bacteria.
Practical – isolating microbes from root nodules
Learning objectives
To show the role of microbes in the nitrogen cycle, how microbes can be grown from root nodules and an example of symbiosis.
Materials
- Plant with root nodules, e.g. clover, peas
- Mannitol yeast extract agar plate (Recipe: Suspend 10 g agar in 1 litre water. Heat to dissolve. Add 0.5 g K2HPO4, 0.2 g MgSO4.7H2O, 0.2 g NaCl, 0.2 g CaCl2.6H2O, 10 g mannitol and 0.4 g yeast extract. Dispense and sterilize by autoclaving.)
- Sterile distilled water in beaker (covered)
- 70% (v/v) industrial denatured alcohol in a small beaker or glass Petri dish covered in foil (caution: flammable, keep covered and away from lit Bunsen burner)
- 5 sterile Petri dishes
- Bunsen burner
- Pasteur pipettes: 1 sterile, 1 non-sterile
- Sterile glass rod
- Scalpel
- Metal forceps (can be pre-sterilized by autoclaving)
- Wire loop
- Beaker of disinfectant
- Discard pot
- Marker pen
- Adhesive tape
Recommendations
- Advance planning is necessary to obtain suitable plant material. Clover (obtain seeds from school science suppliers or plants from a field or lawn) is recommended as the nodules are relatively soft.
- Potato dextrose agar supplemented with 0.25 g yeast extract per litre may be used instead of mannitol yeast extract agar.
- The plates should be incubated preferably at 20–25°C or at room temperature for 2–3 days.
- If the nodules are cleaned well in the alcohol, a population of predominantly Rhizobium should result. Students should be informed that they are using sterile apparatus so that any bacteria that do grow on their plates are likely to have come from the root nodules.
- 1% bleach solution can be used instead of alcohol to clean the nodules.
- The colonies of Rhizobium are off-white with a sticky appearance. Colonies of other colours are not Rhizobium; they may be either intracellular contaminants from the nodule or soil microbes that have survived the washing and alcohol treatment.
Procedure
- Choose a length of root that has nodules and cut off a portion about 1 cm long using a scalpel. Hold the portion of root by forceps and wash free of soil using tap water.
- Transfer several drops of 70% (v/v) industrial denatured alcohol by Pasteur pipette fitted with a teat to a sterile Petri dish. The pipette need not be sterile for this operation; put the pipette into a discard pot. Transfer the washed portion of the root to the alcohol in the Petri dish with forceps and leave immersed for 1–2 minutes to sterilize it. Use aseptic technique from this stage forward.
- Transfer sufficient sterile water to cover the base of another Petri dish using a sterile Pasteur pipette fitted with a teat. If it is necessary to re-use the pipette, keep it sterile, e.g. by resting under the lid of a sterile Petri dish base.
- Use sterile forceps or sterilize them by dipping in alcohol (keeping the points facing downward) and passing quickly through the Bunsen burner flame, allow to cool and use to transfer the portion of root to the sterile water in the Petri dish to rinse off the alcohol. Repeat this operation at least twice more with fresh sterile water. If using alcohol take care to keep the pot well away from the Bunsen burner flame.
- Transfer a few drops of sterile water to a sterile Petri dish and add the portion of root using sterile metal forceps. Macerate the nodules using a sterile glass rod (or forceps) to produce a milky fluid.
- Label the base of a mannitol yeast extract agar plate with your name, the date and MYEA. Sterilize a wire loop by flaming, cool it, take a loopful of the nodule macerate and streak it out on the plate as shown here:
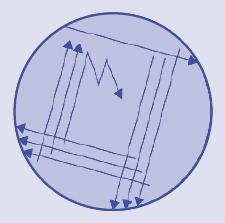
Reflame the loop. Tape the lid on the plate, invert it and incubate for 3–4 days. Dispose of contaminated materials appropriately.
Next lesson...
Examine the MYEA plate without removing the lid, noting the appearance of any colonies growing on the agar.
Reproduced from our education resource, Practical Microbiology for Secondary Schools. This publication and its companion Basic Practical Microbiology (which contains details of how to carry out a risk assessment) are available free to School Members. Please contact [email protected].
