Corynebacterium diphtheriae cares not for our indifference
My name is Dr Michael Weigand and I’m a bioinformatics health scientist within the Division of Bacterial Diseases at the U.S. Centers for Disease Control and Prevention (CDC). Much of my work focuses on genomic analyses of bacterial agents of vaccine-preventable respiratory diseases, notably pertussis and diphtheria. I’m especially focused on utilising methods of advanced molecular detection in pursuit of our agency’s mission to protect people from health threats.
One of those pathogens is Corynebacterium diphtheriae, an aerobic, Gram-positive bacteria, some strains of which produce the deadly diphtheria toxin (‘toxigenic’). As the name implies, the eponymous toxin is responsible for many of the symptoms and complications associated with the severe respiratory disease known as ‘diphtheria.’ The foundation of treatment for diphtheria is application of neutralising antitoxin. Outcomes vary based on healthcare access, but fatality rates may reach 10% during outbreaks and as high as 50% in unvaccinated populations, particularly if antitoxin treatment is unavailable.
Thankfully, effective vaccines against diphtheria have existed for more than 100 years and work by targeting the diphtheria toxin such that cases remain rare in most of the world. However, toxigenic strains of C. diphtheriae are endemic to many regions and outbreaks are more common in places with limitations or disruptions to health care and low vaccination rates. For example, a few years ago our CDC laboratory and epidemiology teams were called upon to assist with investigation of a potential diphtheria outbreak in Bangladesh among forcibly displaced Myanmar people densely settled there in refugee camps. Upon arrival, our team provided diagnostic laboratory support, confirming cases of diphtheria caused by toxigenic strains of C. diphtheriae, and subsequent assistance to implement necessary control measures.
With the immediate public health support tasks completed, we then coordinated with the local ministry of health to ship a selection of cultured isolates back to our laboratory in Atlanta for further characterisation through whole-genome sequencing (WGS). High-throughput sequencing and bioinformatics can reproducibly characterize genomes of C. diphtheriae to evaluate isolate relatedness through multi-locus sequence typing (MLST) and quantifying single nucleotide polymorphisms (SNPs). The genomic data clearly shows the outbreak was caused not by one strain but concurrent transmission following multiple introductions of toxigenic C. diphtheriae, each distinguishable by MLST and phylogenetic reconstruction. These results further emphasised that conditions in the host population, specifically limited health care and low vaccination rates, fuelled the rapid spread of disease. Plus, multiple strains of toxigenic C. diphtheriae were present, ready to seize the opportunity.
To our surprise, the study also identified the presence of non-toxigenic C. diphtheriae in two diphtheria case patients. At this point, I should probably mention genes for diphtheria toxin are encoded on a mobile genetic element known as a ‘corynephage,’ which is not part of the core, genetic backbone. C. diphtheriae lives quite happily without the toxin genes and maintains a toolbox of other virulence factors that help it colonise human hosts. The isolate in our study was recovered from a throat swab, but non-toxigenic C. diphtheriae usually causes cutaneous infection characterized by non-healing and non-distinctive shallow ulcers. Although clinical outcomes are much less severe than toxin-mediated respiratory disease, non-toxigenic strains appear readily transmissible. Resulting cutaneous infection from non-toxigenic strains is more common among persons experiencing homelessness or those who inject drugs. Because these strains do not produce toxin, diphtheria vaccination does not offer protection.
In a previous study, we compared genomic data from non-toxigenic isolates and observed a cluster of patients visiting homeless shelters with non-healing wounds caused by a common strain of C. diphtheriae (Xiaoli et al 2020). It’s striking to me that phylogenetic trees constructed from clusters of toxigenic and non-toxigenic C. diphtheriae infection are indistinguishable underscoring the importance of core virulence factors, not the diphtheria toxin, in host transmission and colonisation. In both contexts, only WGS provides sufficient resolution to infer whether contemporaneous cases may be caused by a rapidly spreading outbreak strain or the sustained transmission of an endemic population.
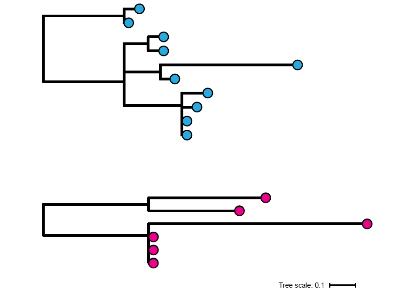
Now back to the diphtheria outbreak in Bangladesh, where we observed two individuals coinfected with divergent strains of toxigenic and non-toxigenic C. diphtheriae. Additional microbiological testing and long-read WGS on the PacBio platform revealed one non-toxigenic strain carried a plasmid with multiple genes conferring drug resistance. Reduced susceptibility of microbial pathogens to antibiotics is a critical threat to public health, and it’s not hard to imagine the potential implications of these two genotypes interacting to transfer this resistance plasmid to the toxigenic strain. Let’s also not forget, the non-toxigenic strain already carrying the plasmid is just one mobile element away from becoming a deadly, toxigenic C. diphtheriae itself. Genetic exchange in either direction could have significant consequences for the control of diphtheria.
All of which is to say, C. diphtheriae—toxigenic or not—is a successful pathogen that circulates widely, even endemic to some regions, awaiting human-made opportunities to cause infection. Toxin-mediated disease rightly deserves the most attention given its severity, but we cannot ignore the public health problems presented by non-toxigenic C. diphtheriae. Circulating non-toxigenic strains place an immediate health burden on marginalised at-risk groups as well as a long-term threat of providing a genetic reservoir for deadly toxigenic strains. Genomic analyses like ours serve as a reminder that a common pathogen is behind the two routes of infection and phylogenetic patterns of their transmission are not that different.
You can read Dr Michael Weigand's othe work published in Microbial Genomics here.
Image: CDC
