The bacteriophage - bacteria’s worst enemy?
Posted on July 3, 2019 by Matt Bassett
Viruses are defined as infective agents that can only multiply within the living cells of a host. Their evolutionary origins are slightly unclear, but theories include evolution from plasmids or bacteria. Viruses can infect all forms of life, and 6 years after their discovery by Dmitri Ivanovsky in 1892, bacteriologist Ernest Hanbury Hankin reported that an unknown agent in the Ganges and Yamuna rivers could kill cholera.

It wasn’t until 1915 that the area had another breakthrough, with the agent being discovered independently by British bacteriologist Frederick Twort in 1915, and by French microbiologist Félix d’Hérelle in 1917. It was realised that these new agents were viruses that parasitized bacteria and d’Hérelle gave these agents the name bacteriophage (commonly shortened to phage), translating as bacteria eater.
Bacteriophages are composed of proteins and a DNA or RNA genome that can be very simple, containing four genes, or complex, with hundreds of genes. The phages infect by injecting their genome into the bacteria which disrupts the bacteria’s normal replication cycle. This makes the bacteria’s ribosomes replicate the viral genome and even creates proteins to assemble the new viruses. The replicated viruses are then released, often by lysis where the cell bursts open, and are free to infect new bacteria.
Marine Recyclers
Since the first reporting of bacteriophages it’s been discovered that they are amongst the most common and diverse entities on the planet. It is estimated that there could be more than 1031 phages, more than every other organism on the planet combined. Phages are found everywhere, but the oceans are truly their home, with researchers estimating there are around 5x107 (50,000,000) phages per millilitre of water. These marine bacteriophages appear to be crucial for the ecology of oceans. When the phages burst out of their host cell through lysis, the organic matter and nutrients within the bacteria are released into the ocean, fulfilling a crucial step within the nutrient cycle known as the viral shunt. In this way marine bacteriophages are thought to cycle carbon through the food web, and also regulate microbial biodiversity by preventing bacterial population explosions.
Phage Therapy
On discovering bacteriophages, Felix d’Hérelle noted that phages always appeared in the stools of patients suffering with Shigella just prior to recovery. This discovery led to increased research into the use of phages against bacterial infections in what became known as phage therapy. In comparison to using antibiotics as medicines, using phages does have benefits. Antibiotics kill bacteria indiscriminately, wiping out beneficial bacteria as well as the pathogens, whereas phages are much more specific as they only kill the pathogenic bacteria and are typically harmless to the host and their gut flora. In addition, bacteriophages have a high therapeutic index, the therapeutic index of a medicine is the measure of how safe a drug is, with a high index meaning they would be expected to have little to no side effects.
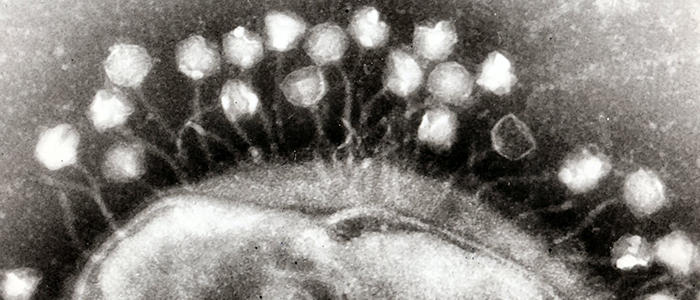
When first discovered there was considerable excitement around bacteriophages for their possible use as medicines in what has become known as phage therapy. However, although the high specificity of phages is a benefit, there are also huge obstacles as vast ‘banks’ of phages are needed and must be regularly updated to keep up with the constantly evolving phages. These constantly changing banks make it difficult and expensive to safely check each treatment. As antibiotics were more effective and less expensive to produce, phage therapy was quickly side-lined. However, with the emerging threat of antimicrobial resistance, alternatives to antibiotics are needed, could phage therapy be considered a viable option?
The Journal of General Virology has launched the Bacteriophage Collection. Guest-edited by Professor Tetsuya Hayashi (Kyushu University, Japan), this collection brings together original research articles, methods, mini and full-length reviews relating to the diversity of bacteriophages and genomics-based research with a focus on their roles in the evolution of bacteria and ecosystems.
If you would like to submit to the collection you can do so here. Please state that your manuscript is intended for the Bacteriophage Collection upon submission.
Further reading from the collection
- Applying phylogenomics to understand the emergence of Shiga-toxin-producing Escherichia coli O157:H7 strains causing severe human disease in the UK
- PhagePhisher: a pipeline for the discovery of covert viral sequences in complex genomic datasets
- Evolution of a zoonotic pathogen: investigating prophage diversity in enterohaemorrhagic Escherichia coli O157 by long-read sequencing