Discovering a new bacteriophage against multidrug-resistant E. coli

Urinary tract infections (UTIs) are among the most common bacterial infections worldwide, affecting millions of people every year. Although many cases are mild and respond well to antibiotics, the rise of multidrug-resistant (MDR) Escherichia coli isolates has made treatment increasingly difficult. As a microbiologist working in Chile, I often encounter clinical strains that resist several families of antibiotics, leaving very limited therapeutic options. This scenario inspired our research group to explore alternative antimicrobial strategies, including bacteriophages, viruses that specifically infect bacteria and, in some cases, can lyse strains that are otherwise untreatable.
My name is Boris Parra, and I am an academic researcher at Universidad de Las Américas (UDLA) and Universidad de Concepción (UDEC). For several years, my work has focused on the interaction between phages and pathogenic bacteria, with a special interest in isolating viruses capable of targeting MDR microbes relevant to public health. Our recently published paper describes the discovery and characterization of EcoPhCCP1, a novel lytic bacteriophage able to infect multiple MDR E. coli isolates recovered from urinary tract infections.
The project began with a simple but persistent question: Could naturally occurring phages be used to target the MDR strains that clinicians struggle to treat? UTIs represent a major healthcare burden in Chile and globally, and the increasing prevalence of extended-spectrum β-lactamase (ESBL) producers and carbapenem-resistant E. coli complicates patient management even further. We believed that searching for phages in environmental samples—water sources, sewage, and urban environments—could reveal viruses naturally adapted to infect these difficult strains.
Our initial screening involved enriching environmental samples with clinically relevant E. coli isolates. Very early in the process, one enrichment produced an unusually clear and large zone of lysis, suggesting the presence of a potent lytic phage. That was our first encounter with EcoPhCCP1. Even before purification, the plaques it generated hinted at a virus capable of rapid and efficient replication. Those plaques, simple as they were, quickly became a source of excitement in the lab: could this be the phage we were hoping to find?
Once isolated, EcoPhCCP1 demonstrated a surprisingly broad host range for an E. coli phage. It was able to lyse multiple MDR strains obtained from UTI cases, including ESBL producers and isolates resistant to fluoroquinolones and aminoglycosides. This broad activity suggested a phage with valuable therapeutic potential, at least at the experimental level. We then moved forward with a deeper biological and genomic characterization.
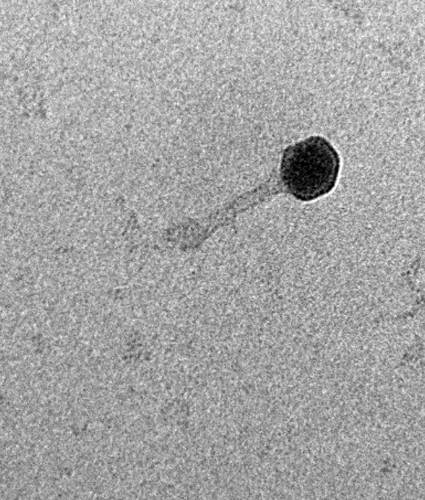
Electron microscopy revealed that EcoPhCCP1 belongs to the Caudoviricetes class, displaying the morphology typical of lytic tailed phages. Its infection cycle was fast, with short latent periods and high burst sizes, features commonly associated with efficient bacterial killing. Still, morphology and infection dynamics were only part of the story—genomics was essential to establish safety, taxonomy, and evolutionary context.
Whole-genome sequencing showed that EcoPhCCP1 carries a linear dsDNA genome with hallmark structural and replication proteins consistent with strictly lytic phages. Importantly, the genome lacked genes associated with lysogeny, antibiotic resistance, or bacterial virulence. These findings support the potential safety of EcoPhCCP1 for future applications, at least within an experimental or preclinical framework. Genomic comparison also revealed that EcoPhCCP1 shares limited similarity with previously described phages, suggesting it may represent a new species within its genus.
Studying the genome was a particularly memorable stage of the project. There is something uniquely satisfying about identifying each gene, predicting its function, and seeing how the molecular architecture of the virus explains the behavior we observed experimentally. It was also motivating to watch undergraduate students in our group take ownership of different steps—plaque assays, DNA extractions, host-range tests—and grow scientifically throughout the project. Their contributions were essential, and the excitement of seeing the genome assembled for the first time is something we all shared.
Another important part of the story was connecting the biological data with the broader problem we hoped to address. MDR UTIs cause recurrent infections, prolonged hospital stays, and high antibiotic consumption. Many clinicians are forced to rely on last-line antibiotics such as carbapenems or fosfomycin. In this context, an effective lytic phage is not just a scientific curiosity; it represents a potential tool to complement existing treatments or inspire new therapeutic strategies. While we are still far from clinical use, the foundational characterization of promising phages is a necessary first step.
Looking ahead, we hope to expand this line of research by testing EcoPhCCP1 in more complex infection models, exploring its stability under conditions relevant to phage therapy, and evaluating possible synergistic interactions with antibiotics. We are also interested in identifying additional phages with complementary host ranges, which could eventually form the basis of a rationally designed phage cocktail for MDR E. coli. The field of phage therapy is growing rapidly worldwide, and countries in Latin America—including Chile—are increasingly recognizing the importance of developing local expertise.
Why should we be excited about this research area? Because phages offer something unique: precision. Unlike broad-spectrum antibiotics, they can selectively eliminate pathogenic bacteria while sparing the surrounding microbiota. Their diversity is immense, and the natural world still holds an enormous number of phages waiting to be discovered. Each new virus provides an opportunity to better understand microbial ecosystems and to develop alternatives to antibiotics at a time when antimicrobial resistance poses one of the greatest global health challenges.
EcoPhCCP1 is only one of many phages that remain hidden in the environment, ready to be found. Our work highlights how classical microbiology, combined with modern genomics, can reveal new biological tools with real-world potential. The journey behind this phage—from a water sample to a fully sequenced and characterized virus—showcases how curiosity-driven research can intersect with pressing clinical problems. And it reminds us that the smallest biological entities can sometimes hold the biggest promise.